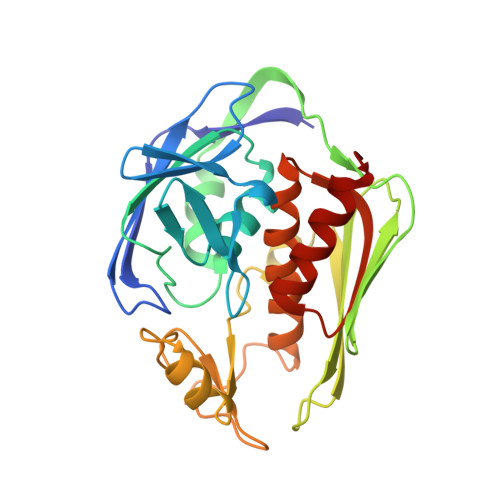
Binding of Uridine 5-Diphosphate in the Basic Patch of the Zinc Metalloenzyme Deacetylase LpxC and Implications for Substrate Binding
Gennadios, H.A., Christianson, D.W.(2006) Biochemistry 45: 15216-15223
- PubMed: 17176043
- DOI: https://doi.org/10.1021/bi0619021
- Primary Citation of Related Structures:
2IER, 2IES - PubMed Abstract:
LpxC is a zinc metalloenzyme that catalyzes the first committed step in the biosynthesis of lipid A, a vital component of the outer membrane of Gram-negative bacteria. Accordingly, the inhibition of LpxC is an attractive strategy for the treatment of Gram-negative bacterial infections. Here, we report the 2.7 A resolution X-ray crystal structure of LpxC from Aquifex aeolicus complexed with uridine 5'-diphosphate (UDP), and the 3.1 A resolution structure of LpxC complexed with pyrophosphate. The X-ray crystal structure of the LpxC-UDP complex provides the first view of interactions likely to be exploited by the substrate UDP group in the "basic patch" of the active site. The diphosphate group of UDP makes hydrogen bond interactions with strictly conserved residue K239 as well as solvent molecules. The ribose moiety of UDP interacts with partially conserved residue E197. The UDP uracil group hydrogen bonds with both the backbone NH group and the backbone carbonyl group of E160, and with the backbone NH group of K162 through an intervening water molecule. Finally, the alpha-phosphate and uracil groups of UDP interact with R143 and R262 through intervening water molecules. The structure of LpxC complexed with pyrophosphate reveals generally similar intermolecular interactions in the basic patch. Unexpectedly, diphosphate binding in both complexes is accompanied by coordination to an additional zinc ion, resulting in the identification of a new metal-binding site termed the E-site. The structures of the LpxC-UDP and LpxC-pyrophosphate complexes provide new insights with regard to substrate recognition in the basic patch and metal ion coordination in the active site of LpxC.
Organizational Affiliation:
Roy and Diana Vagelos Laboratories, Department of Chemistry, University of Pennsylvania, Philadelphia, Pennsylvania 19104-6323.