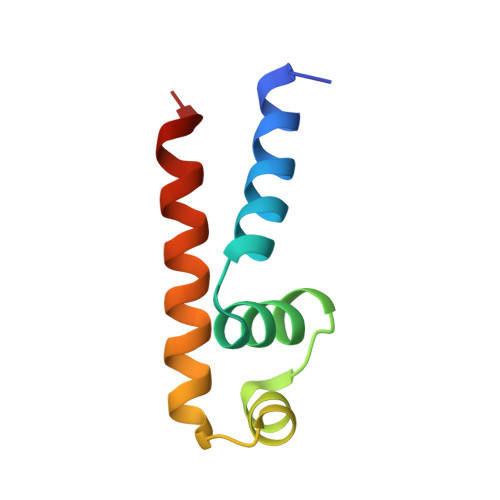
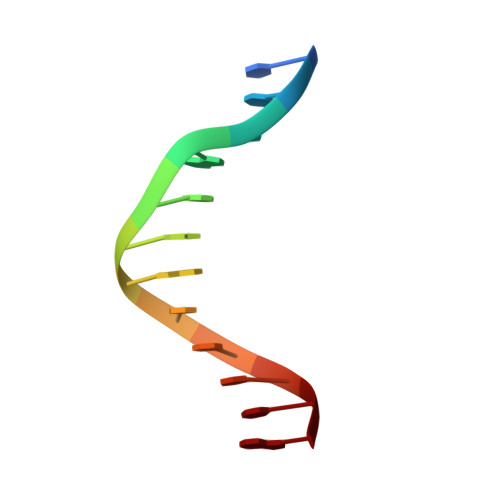
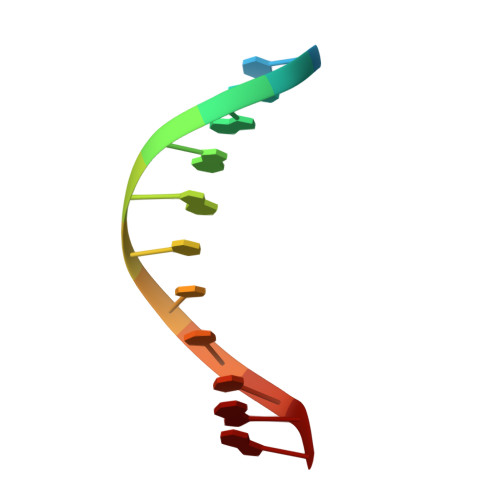
The structural basis for promoter -35 element recognition by the group IV sigma factors.
Lane, W.J., Darst, S.A.(2006) PLoS Biol 4: e269-e269
- PubMed: 16903784
- DOI: https://doi.org/10.1371/journal.pbio.0040269
- Primary Citation of Related Structures:
2H27 - PubMed Abstract:
The control of bacterial transcription initiation depends on a primary sigma factor for housekeeping functions, as well as alternative sigma factors that control regulons in response to environmental stresses. The largest and most diverse subgroup of alternative sigma factors, the group IV extracytoplasmic function sigma factors, directs the transcription of genes that regulate a wide variety of responses, including envelope stress and pathogenesis. We determined the 2.3-A resolution crystal structure of the -35 element recognition domain of a group IV sigma factor, Escherichia coli sigma(E)4, bound to its consensus -35 element, GGAACTT. Despite similar function and secondary structure, the primary and group IV sigma factors recognize their -35 elements using distinct mechanisms. Conserved sequence elements of the sigma(E) -35 element induce a DNA geometry characteristic of AA/TT-tract DNA, including a rigid, straight double-helical axis and a narrow minor groove. For this reason, the highly conserved AA in the middle of the GGAACTT motif is essential for -35 element recognition by sigma(E)4, despite the absence of direct protein-DNA interactions with these DNA bases. These principles of sigma(E)4/-35 element recognition can be applied to a wide range of other group IV sigma factors.
Organizational Affiliation:
The Rockefeller University, New York, New York, United States of America.