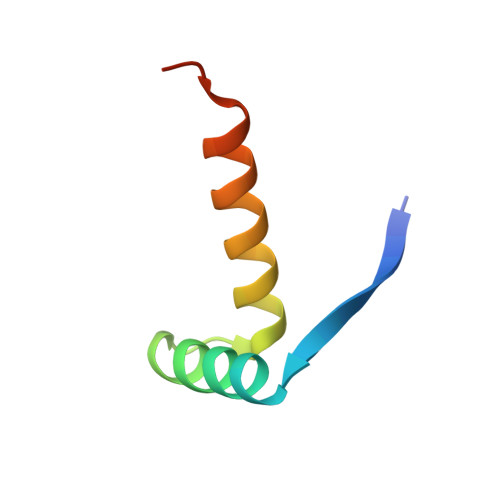
Crystal structures of the DNA-binding domain of Escherichia coli proline utilization A flavoprotein and analysis of the role of Lys9 in DNA recognition.
Larson, J.D., Jenkins, J.L., Schuermann, J.P., Zhou, Y., Becker, D.F., Tanner, J.J.(2006) Protein Sci 15: 2630-2641
- PubMed: 17001030
- DOI: https://doi.org/10.1110/ps.062425706
- Primary Citation of Related Structures:
2AY0, 2GPE - PubMed Abstract:
PutA (proline utilization A) from Escherichia coli is a 1320-amino-acid residue protein that is both a bifunctional proline catabolic enzyme and an autogenous transcriptional repressor. Here, we report the first crystal structure of a PutA DNA-binding domain along with functional analysis of a mutant PutA defective in DNA binding. Crystals were grown using a polypeptide corresponding to residues 1-52 of E. coli PutA (PutA52). The 2.1 Angstrom resolution structure of PutA52 mutant Lys9Met was determined using Se-Met MAD phasing, and the structure of native PutA52 was solved at 1.9 Angstrom resolution using molecular replacement. Residues 3-46 form a ribbon-helix-helix (RHH) substructure, thus establishing PutA as the largest protein to contain an RHH domain. The PutA RHH domain forms the intertwined dimer with tightly packed hydrophobic core that is characteristic of the RHH family. The structures were used to examine the three-dimensional context of residues conserved in PutA RHH domains. Homology modeling suggests that Lys9 and Thr5 contact DNA bases through the major groove, while Arg15, Thr28, and His30 may interact with the phosphate backbone. Lys9 is shown to be essential for specific recognition of put control DNA using gel shift analysis of the Lys9Met mutant of full-length PutA. Lys9 is disordered in the PutA52 structure, which implies an induced-fit binding mechanism in which the side chain of Lys9 becomes ordered through interaction with DNA. These results provide new insights into the structural basis of DNA recognition by PutA and reveal three-dimensional structural details of the PutA dimer interface.
- Department of Chemistry, University of Missouri--Columbia, Columbia, Missouri 65211, USA.
Organizational Affiliation: