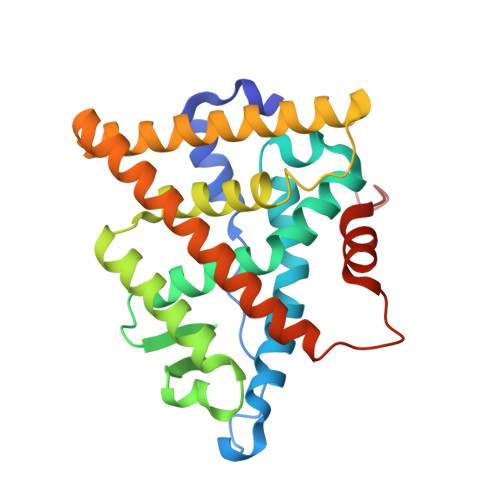
The 2.0 A crystal structure of the ER{alpha} ligand-binding domain complexed with lasofoxifene
Vajdos, F.F., Hoth, L.R., Geoghegan, K.F., Simons, S.P., LeMotte, P.K., Danley, D.E., Ammirati, M.J., Pandit, J.(2007) Protein Sci 16: 897-905
- PubMed: 17456742
- DOI: https://doi.org/10.1110/ps.062729207
- Primary Citation of Related Structures:
2OUZ - PubMed Abstract:
Lasofoxifene is a new and potent selective estrogen receptor modulator (SERM). The structural basis of its interaction with the estrogen receptor has been investigated by crystallographic analysis of its complex with the ligand-binding domain of estrogen receptor alpha at a resolution of 2.0 A. As with other SERMs, lasofoxifene diverts the receptor from its agonist-bound conformation by displacing the C-terminal AF-2 helix into the site at which the LXXLL motif of coactivator proteins would otherwise be able to bind. Lasofoxifene achieves this effect by occupying the space normally filled by residue Leu 540, as well as by modulating the conformation of residues of helix 11 (His 524, Leu 525). A well-defined salt bridge between lasofoxifene and Asp 351 suggests that charge neutralization in this region of the receptor may explain the some of the antiestrogenic effects of lasofoxifene. The results suggest general features of ERalpha/SERM recognition, and add a new dimension to efforts to rationalize differences between the biological activity profiles exhibited by these important pharmacological agents.
Organizational Affiliation:
Department of Exploratory Medicinal Sciences, Pfizer Global Research and Development, Pfizer Inc., Groton, Connecticut 06340-8001, USA. felix.vajdos@pfizer.com