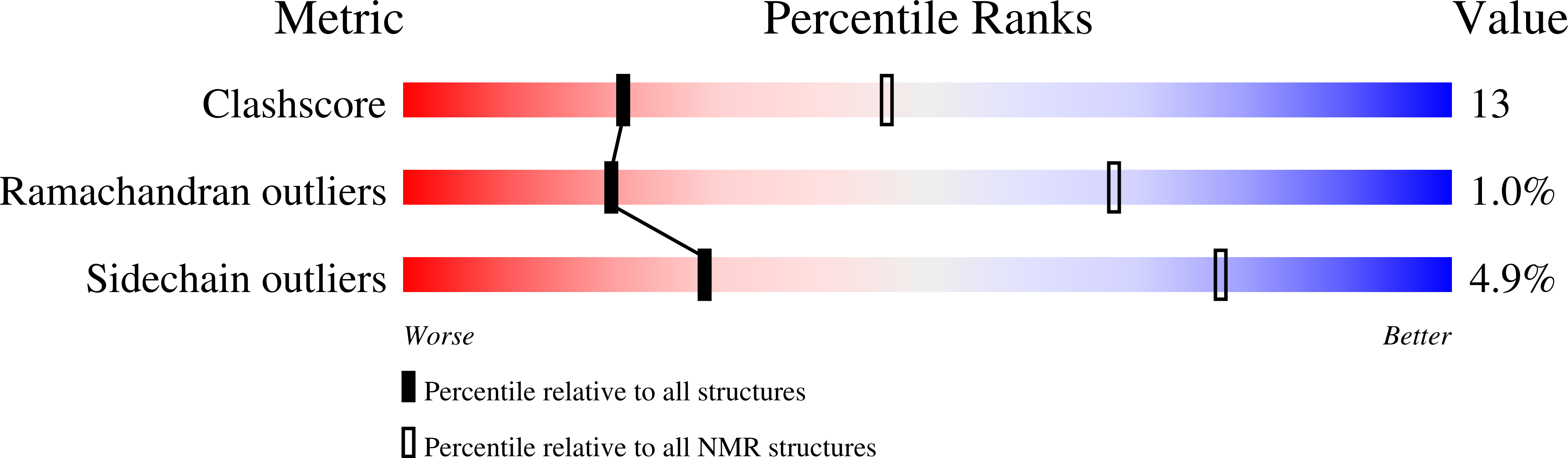Engineering Metamorphic Chemokine Lymphotactin/XCL1 into the GAG-Binding, HIV-Inhibitory Dimer Conformation.
Fox, J.C., Tyler, R.C., Guzzo, C., Tuinstra, R.L., Peterson, F.C., Lusso, P., Volkman, B.F.(2015) ACS Chem Biol 10: 2580-2588
- PubMed: 26302421
- DOI: https://doi.org/10.1021/acschembio.5b00542
- Primary Citation of Related Structures:
2N54 - PubMed Abstract:
Unlike other chemokines, XCL1 undergoes a distinct metamorphic interconversion between a canonical monomeric chemokine fold and a unique β-sandwich dimer. The monomeric conformation binds and activates the receptor XCR1, whereas the dimer binds extracellular matrix glycosaminoglycans and has been associated with anti-human immunodeficiency virus (HIV) activity. Functional studies of WT-XCL1 are complex, as both conformations are populated in solution. To overcome this limitation, we engineered a stabilized dimeric variant of XCL1 designated CC5. This variant features a new disulfide bond (A36C-A49C) that prevents structural interconversion by locking the chemokine into the β-sandwich dimeric conformation, as demonstrated by NMR structural analysis and hydrogen/deuterium exchange experiments. Functional studies analyzing glycosaminoglycan binding demonstrate that CC5 binds with high affinity to heparin. In addition, CC5 exhibits potent inhibition of HIV-1 activity in primary peripheral blood mononuclear cells (PBMCs), demonstrating the importance of the dimer in blocking viral infection. Conformational variants like CC5 are valuable tools for elucidating the biological relevance of the XCL1 native-state interconversion and will assist in future antiviral and functional studies.
Organizational Affiliation:
Department of Biochemistry, Medical College of Wisconsin , Milwaukee, Wisconsin 53226, United States.