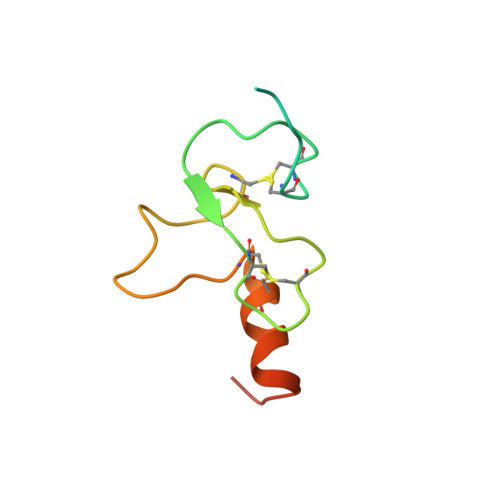
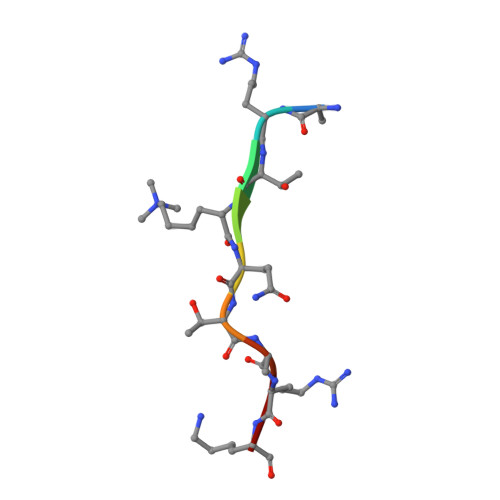
Yng1 PHD finger binding to H3 trimethylated at K4 promotes NuA3 HAT activity at K14 of H3 and transcription at a subset of targeted ORFs
Taverna, S.D., Ilin, S., Rogers, R.S., Tanny, J.C., Lavender, H., Li, H., Baker, L., Boyle, J., Blair, L.P., Chait, B.T., Patel, D.J., Aitchison, J.D., Tackett, A.J., Allis, C.D.(2006) Mol Cell 24: 785-796
- PubMed: 17157260
- DOI: https://doi.org/10.1016/j.molcel.2006.10.026
- Primary Citation of Related Structures:
2JMI, 2JMJ - PubMed Abstract:
Posttranslational histone modifications participate in modulating the structure and function of chromatin. Promoters of transcribed genes are enriched with K4 trimethylation and hyperacetylation on the N-terminal tail of histone H3. Recently, PHD finger proteins, like Yng1 in the NuA3 HAT complex, were shown to interact with H3K4me3, indicating a biochemical link between K4 methylation and hyperacetylation. By using a combination of mass spectrometry, biochemistry, and NMR, we detail the Yng1 PHD-H3K4me3 interaction and the importance of NuA3-dependent acetylation at K14. Furthermore, genome-wide ChIP-Chip analysis demonstrates colocalization of Yng1 and H3K4me3 in vivo. Disrupting the K4me3 binding of Yng1 altered K14ac and transcription at certain genes, thereby demonstrating direct in vivo evidence of sequential trimethyl binding, acetyltransferase activity, and gene regulation by NuA3. Our data support a general mechanism of transcriptional control through which histone acetylation upstream of gene activation is promoted partially through availability of H3K4me3, "read" by binding modules in select subunits.
Organizational Affiliation:
Laboratory of Chromatin Biology, The Rockefeller University.