Biochemical and structural basis for partially redundant enzymatic and transcriptional functions of DCoH and DCoH2
Rose, R.B., Pullen, K.E., Bayle, J.H., Crabtree, G.R., Alber, T.(2004) Biochemistry 43: 7345-7355
- PubMed: 15182178
- DOI: https://doi.org/10.1021/bi049620t
- Primary Citation of Related Structures:
1RU0 - PubMed Abstract:
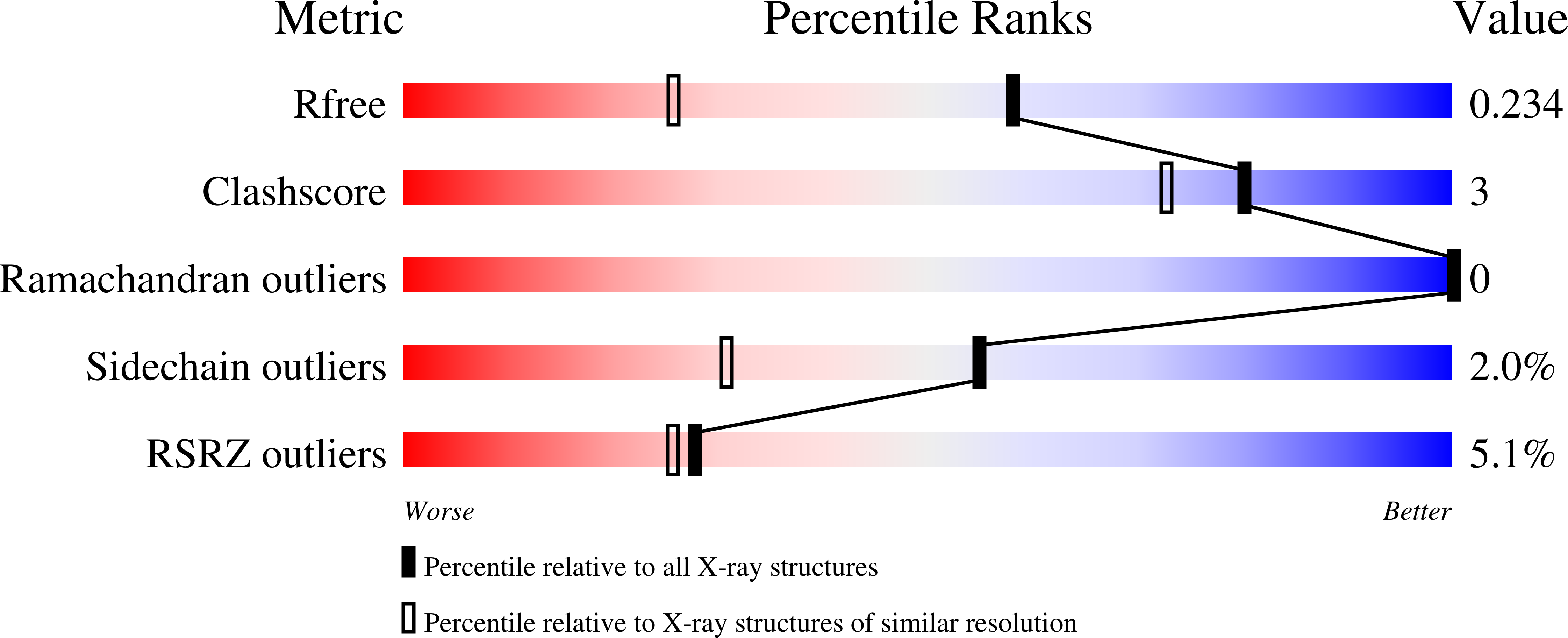
An inherited form of diabetes, maturity-onset diabetes of the young type 3 (MODY3), results from mutations in the transcriptional activator, hepatocyte nuclear factor-1alpha (HNF1alpha). Transcription by HNF1alpha is stimulated by the bifunctional coactivator DCoH (dimerization cofactor of HNF1). Strikingly, an HNF1alpha deletion in mice causes more severe phenotypes than a DCoH deletion. It has been hypothesized that a DCoH homolog, DCoH2, partially complements the DCoH deletion. To test this idea, we determined the biochemical properties and the 1.6-A-resolution crystal structure of DCoH2. Like DCoH, DCoH2 forms a tetramer, displays pterin-4alpha-carbinolamine dehydratase activity, and binds HNF1alpha in vivo and in vitro. DCoH and DCoH2 adopt identical folds with structural differences confined largely to the protein surfaces and the tetramer interface. In contrast to the hyperstable DCoH tetramer, DCoH2 readily disproportionates and forms a 2:2 complex with HNF1 in vitro. Phylogenetic analysis reveals six major subfamilies of DCoH proteins, including unique DCoH and DCoH2 branches in metazoans. These results suggest distinct roles for DCoH and DCoH2. Differences in conserved surface residues could mediate binding to different effectors. We propose that HNF1alpha binding kinetics may distinguish regulation by DCoH2, under thermodynamic control, from regulation by DCoH, under kinetic control.
Organizational Affiliation:
Department of Molecular and Cell Biology, University of California, Berkeley, California 94720-3206, USA. bob_rose@ncsu.edu