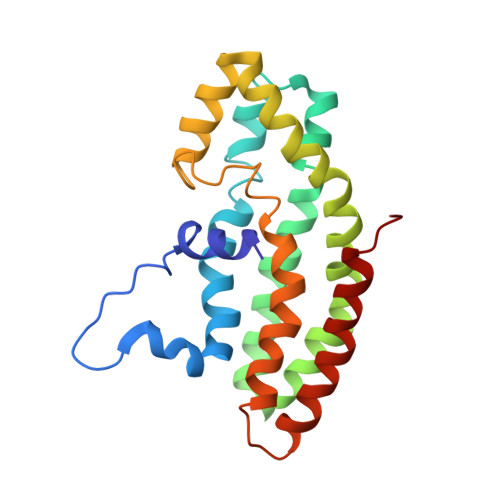
Structural basis for relief of autoinhibition of the Dbl homology domain of proto-oncogene Vav by tyrosine phosphorylation.
Aghazadeh, B., Lowry, W.E., Huang, X.Y., Rosen, M.K.(2000) Cell 102: 625-633
- PubMed: 11007481
- DOI: https://doi.org/10.1016/s0092-8674(00)00085-4
- Primary Citation of Related Structures:
1F5X - PubMed Abstract:
Rho-family GTPases transduce signals from receptors leading to changes in cell shape and motility, mitogenesis, and development. Proteins containing the Dbl homology (DH) domain are responsible for activating Rho GTPases by catalyzing the exchange of GDP for GTP. Receptor-initiated stimulation of Dbl protein Vav exchange activity involves tyrosine phosphorylation. We show through structure determination that the mVav1 DH domain is autoinhibited by an N-terminal extension, which lies in the GTPase interaction site. This extension contains the Tyr174 Src-family kinase recognition site, and phosphorylation or truncation of this peptide results in stimulation of GEF activity. NMR spectroscopy data show that the N-terminal peptide is released from the DH domain and becomes unstructured upon phosphorylation. Thus, tyrosine phosphorylation relieves autoinhibition by exposing the GTPase interaction surface of the DH domain, which is obligatory for Vav activation.
Organizational Affiliation:
Cellular Biochemistry and Biophysics Program, Memorial Sloan-Kettering Cancer Center, New York, New York 10021, USA.