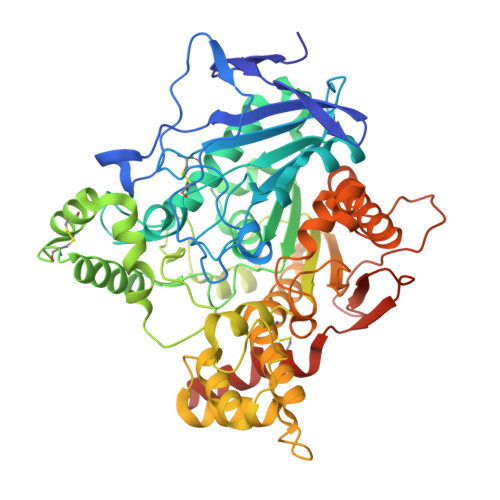
3D Structure of Torpedo Californica Acetylcholinesterase Complexed with Huprine X at 2. 1 A Resolution: Kinetic and Molecular Dynamic Correlates.
Dvir, H., Wong, D.M., Harel, M., Barril, X., Orozco, M., Luque, F.J., Munoz-Torrero, D., Camps, P., Rosenberry, T.L., Silman, I., Sussman, J.L.(2002) Biochemistry 41: 2970
- PubMed: 11863435
- DOI: https://doi.org/10.1021/bi011652i
- Primary Citation of Related Structures:
1E66 - PubMed Abstract:
Huprine X is a novel acetylcholinesterase (AChE) inhibitor, with one of the highest affinities reported for a reversible inhibitor. It is a synthetic hybrid that contains the 4-aminoquinoline substructure of one anti-Alzheimer drug, tacrine, and a carbobicyclic moiety resembling that of another AChE inhibitor, (-)-huperzine A. Cocrystallization of huprine X with Torpedo californica AChE yielded crystals whose 3D structure was determined to 2.1 A resolution. The inhibitor binds to the anionic site and also hinders access to the esteratic site. Its aromatic portion occupies the same binding site as tacrine, stacking between the aromatic rings of Trp84 and Phe330, whereas the carbobicyclic unit occupies the same binding pocket as (-)-huperzine A. Its chlorine substituent was found to lie in a hydrophobic pocket interacting with rings of the aromatic residues Trp432 and Phe330 and with the methyl groups of Met436 and Ile439. Steady-state inhibition data show that huprine X binds to human AChE and Torpedo AChE 28- and 54-fold, respectively, more tightly than tacrine. This difference stems from the fact that the aminoquinoline moiety of huprine X makes interactions similar to those made by tacrine, but additional bonds to the enzyme are made by the huperzine-like substructure and the chlorine atom. Furthermore, both tacrine and huprine X bind more tightly to Torpedo than to human AChE, suggesting that their quinoline substructures interact better with Phe330 than with Tyr337, the corresponding residue in the human AChE structure. Both (-)-huperzine A and huprine X display slow binding properties, but only binding of the former causes a peptide flip of Gly117.
- Department of Structural Biology, Weizmann Institute of Science, Rehovot, Israel, 76100.
Organizational Affiliation: