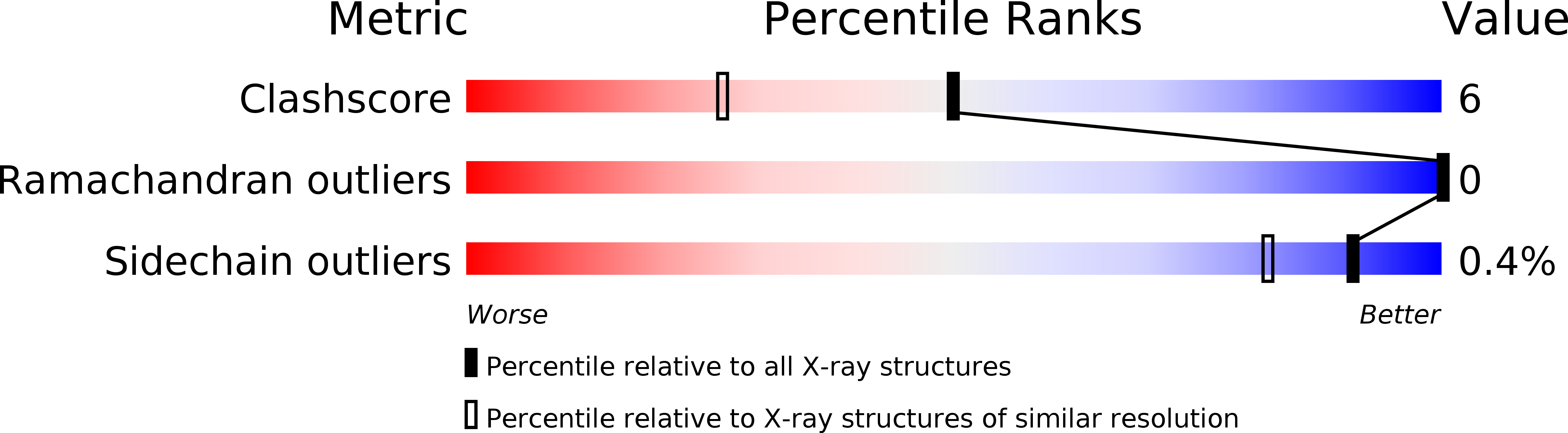The Kappa-Carrageenase of P. Carrageenovora Features a Tunnel-Shaped Active Site: A Novel Insight in the Evolution of Clan-B Glycoside Hydrolases
Michel, G., Chantalat, L., Duee, E., Barbeyron, T., Henrissat, B., Kloareg, B., Dideberg, O.(2001) Structure 9: 513
- PubMed: 11435116
- DOI: https://doi.org/10.1016/s0969-2126(01)00612-8
- Primary Citation of Related Structures:
1DYP - PubMed Abstract:
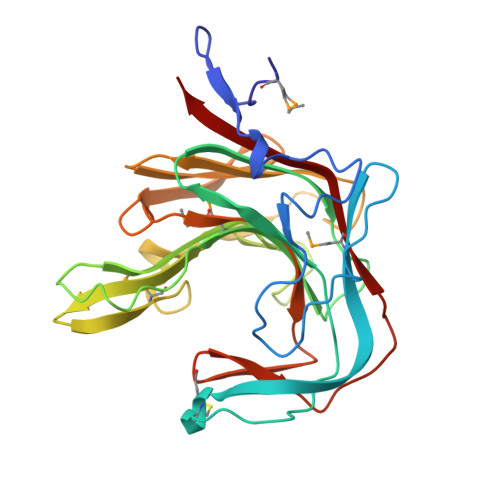
kappa-carrageenans are gel-forming, sulfated 1,3-alpha-1,4-beta-galactans from the cell walls of marine red algae. The kappa-carrageenase from the marine, gram-negative bacterium Pseudoalteromonas carrageenovora degrades kappa-carrageenan both in solution and in solid state by an endoprocessive mechanism. This beta-galactanase belongs to the clan-B of glycoside hydrolases.
Organizational Affiliation:
Laboratoire de Cristallographie Macromoléculaire, Institut de Biologie Structurale Jean-Pierre Ebel, CNRS/CEA, 41 Avenue des Martyrs, 38027 Cedex 1, Grenoble, France.