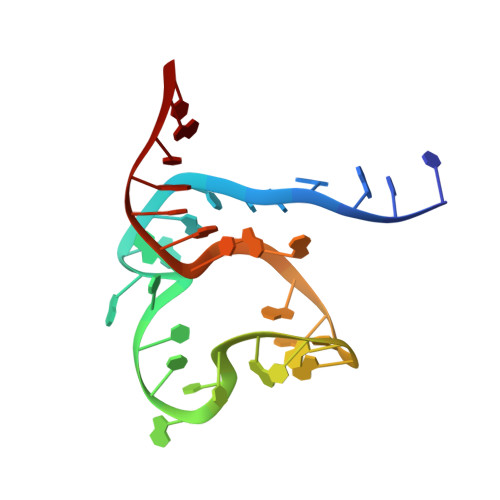
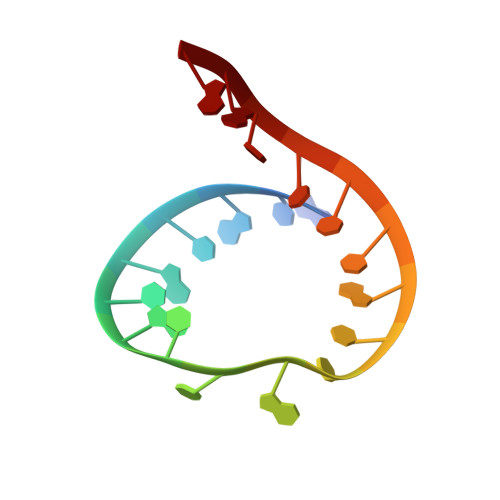
Time-resolved structural analysis of an RNA-cleaving DNA catalyst.
Borggrafe, J., Victor, J., Rosenbach, H., Viegas, A., Gertzen, C.G.W., Wuebben, C., Kovacs, H., Gopalswamy, M., Riesner, D., Steger, G., Schiemann, O., Gohlke, H., Span, I., Etzkorn, M.(2022) Nature 601: 144-149
- PubMed: 34949858
- DOI: https://doi.org/10.1038/s41586-021-04225-4
- Primary Citation of Related Structures:
7PDU - PubMed Abstract:
The 10-23 DNAzyme is one of the most prominent catalytically active DNA sequences 1,2 . Its ability to cleave a wide range of RNA targets with high selectivity entails a substantial therapeutic and biotechnological potential 2 . However, the high expectations have not yet been met, a fact that coincides with the lack of high-resolution and time-resolved information about its mode of action 3 . Here we provide high-resolution NMR characterization of all apparent states of the prototypic 10-23 DNAzyme and present a comprehensive survey of the kinetics and dynamics of its catalytic function. The determined structure and identified metal-ion-binding sites of the precatalytic DNAzyme-RNA complex reveal that the basis of the DNA-mediated catalysis is an interplay among three factors: an unexpected, yet exciting molecular architecture; distinct conformational plasticity; and dynamic modulation by metal ions. We further identify previously hidden rate-limiting transient intermediate states in the DNA-mediated catalytic process via real-time NMR measurements. Using a rationally selected single-atom replacement, we could considerably enhance the performance of the DNAzyme, demonstrating that the acquired knowledge of the molecular structure, its plasticity and the occurrence of long-lived intermediate states constitutes a valuable starting point for the rational design of next-generation DNAzymes.
Organizational Affiliation:
Institut für Physikalische Biologie, Heinrich Heine University Düsseldorf, Düsseldorf, Germany.