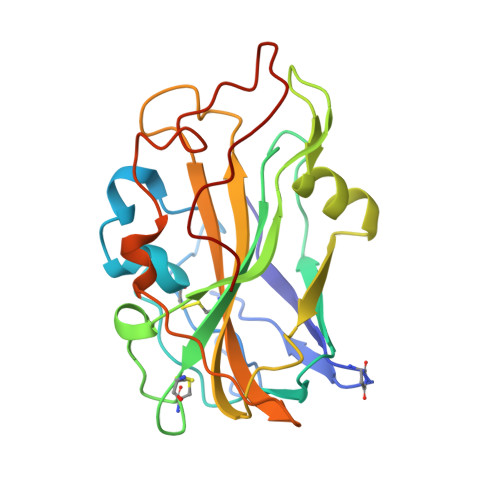
Structure of a C1/C4-oxidizing AA9 lytic polysaccharide monooxygenase from the thermophilic fungus Malbranchea cinnamomea.
Mazurkewich, S., Seveso, A., Huttner, S., Branden, G., Larsbrink, J.(2021) Acta Crystallogr D Struct Biol 77: 1019-1026
- PubMed: 34342275
- DOI: https://doi.org/10.1107/S2059798321006628
- Primary Citation of Related Structures:
7NTL - PubMed Abstract:
The thermophilic fungus Malbranchea cinnamomea contains a host of enzymes that enable its ability as an efficient degrader of plant biomass and that could be mined for industrial applications. This thermophilic fungus has been studied and found to encode eight lytic polysaccharide monooxygenases (LPMOs) from auxiliary activity family 9 (AA9), which collectively possess different substrate specificities for a range of plant cell-wall-related polysaccharides and oligosaccharides. To gain greater insight into the molecular determinants defining the different specificities, structural studies were pursued and the structure of McAA9F was determined. The enzyme contains the immunoglobulin-like fold typical of previously solved AA9 LPMO structures, but contains prominent differences in the loop regions found on the surface of the substrate-binding site. Most significantly, McAA9F has a broad substrate specificity, with activity on both crystalline and soluble polysaccharides. Moreover, it contains a small loop in a region where a large loop has been proposed to govern specificity towards oligosaccharides. The presence of the small loop leads to a considerably flatter and more open surface that is likely to enable the broad specificity of the enzyme. The enzyme contains a succinimide residue substitution, arising from intramolecular cyclization of Asp10, at a position where several homologous members contain an equivalent residue but cyclization has not previously been observed. This first structure of an AA9 LPMO from M. cinnamomea aids both the understanding of this family of enzymes and the exploration of the repertoire of industrially relevant lignocellulolytic enzymes from this fungus.
- Wallenberg Wood Science Center, Division of Industrial Biotechnology, Department of Biology and Biological Engineering, Chalmers University of Technology, SE-412 96 Gothenburg, Sweden.
Organizational Affiliation: