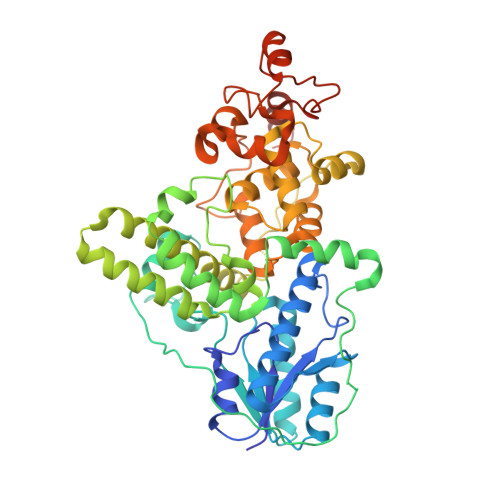
Structural insights into photoactivation of plant Cryptochrome-2.
Palayam, M., Ganapathy, J., Guercio, A.M., Tal, L., Deck, S.L., Shabek, N.(2021) Commun Biol 4: 28-28
- PubMed: 33398020
- DOI: https://doi.org/10.1038/s42003-020-01531-x
- Primary Citation of Related Structures:
6X24 - PubMed Abstract:
Cryptochromes (CRYs) are evolutionarily conserved photoreceptors that mediate various light-induced responses in bacteria, plants, and animals. Plant cryptochromes govern a variety of critical growth and developmental processes including seed germination, flowering time and entrainment of the circadian clock. CRY's photocycle involves reduction of their flavin adenine dinucleotide (FAD)-bound chromophore, which is completely oxidized in the dark and semi to fully reduced in the light signaling-active state. Despite the progress in characterizing cryptochromes, important aspects of their photochemistry, regulation, and light-induced structural changes remain to be addressed. In this study, we determine the crystal structure of the photosensory domain of Arabidopsis CRY2 in a tetrameric active state. Systematic structure-based analyses of photo-activated and inactive plant CRYs elucidate distinct structural elements and critical residues that dynamically partake in photo-induced oligomerization. Our study offers an updated model of CRYs photoactivation mechanism as well as the mode of its regulation by interacting proteins.
Organizational Affiliation:
Department of Plant Biology, University of California - Davis, One shields Avenue, 1002 Life sciences, Davis, CA, 95616, USA.