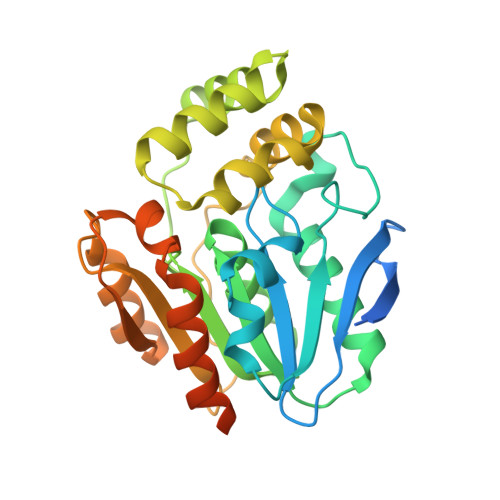
The bottromycin epimerase BotH defines a group of atypical alpha / beta-hydrolase-fold enzymes.
Sikandar, A., Franz, L., Adam, S., Santos-Aberturas, J., Horbal, L., Luzhetskyy, A., Truman, A.W., Kalinina, O.V., Koehnke, J.(2020) Nat Chem Biol 16: 1013-1018
- PubMed: 32601484
- DOI: https://doi.org/10.1038/s41589-020-0569-y
- Primary Citation of Related Structures:
6T6H, 6T6X, 6T6Y, 6T6Z, 6T70 - PubMed Abstract:
D-amino acids endow peptides with diverse, desirable properties, but the post-translational and site-specific epimerization of L-amino acids into their D-counterparts is rare and chemically challenging. Bottromycins are ribosomally synthesized and post-translationally modified peptides that have overcome this challenge and feature a D-aspartate (D-Asp), which was proposed to arise spontaneously during biosynthesis. We have identified the highly unusual α/β-hydrolase (ABH) fold enzyme BotH as a peptide epimerase responsible for the post-translational epimerization of L-Asp to D-Asp during bottromycin biosynthesis. The biochemical characterization of BotH combined with the structures of BotH and the BotH-substrate complex allowed us to propose a mechanism for this reaction. Bioinformatic analyses of BotH homologs show that similar ABH enzymes are found in diverse biosynthetic gene clusters. This places BotH as the founding member of a group of atypical ABH enzymes that may be able to epimerize non-Asp stereocenters across different families of secondary metabolites.
Organizational Affiliation:
Workgroup Structural Biology of Biosynthetic Enzymes, Helmholtz Institute for Pharmaceutical Research Saarland (HIPS), Helmholtz Centre for Infection Research (HZI), Saarland University, Saarbrücken, Germany.