Nek7 conformational flexibility and inhibitor binding probed through protein engineering of the R-spine.
Byrne, M.J., Nasir, N., Basmadjian, C., Bhatia, C., Cunnison, R.F., Carr, K.H., Mas-Droux, C., Yeoh, S., Cano, C., Bayliss, R.(2020) Biochem J 477: 1525-1539
- PubMed: 32242624
- DOI: https://doi.org/10.1042/BCJ20200128
- Primary Citation of Related Structures:
6S73, 6S75, 6S76, 6SK9 - PubMed Abstract:
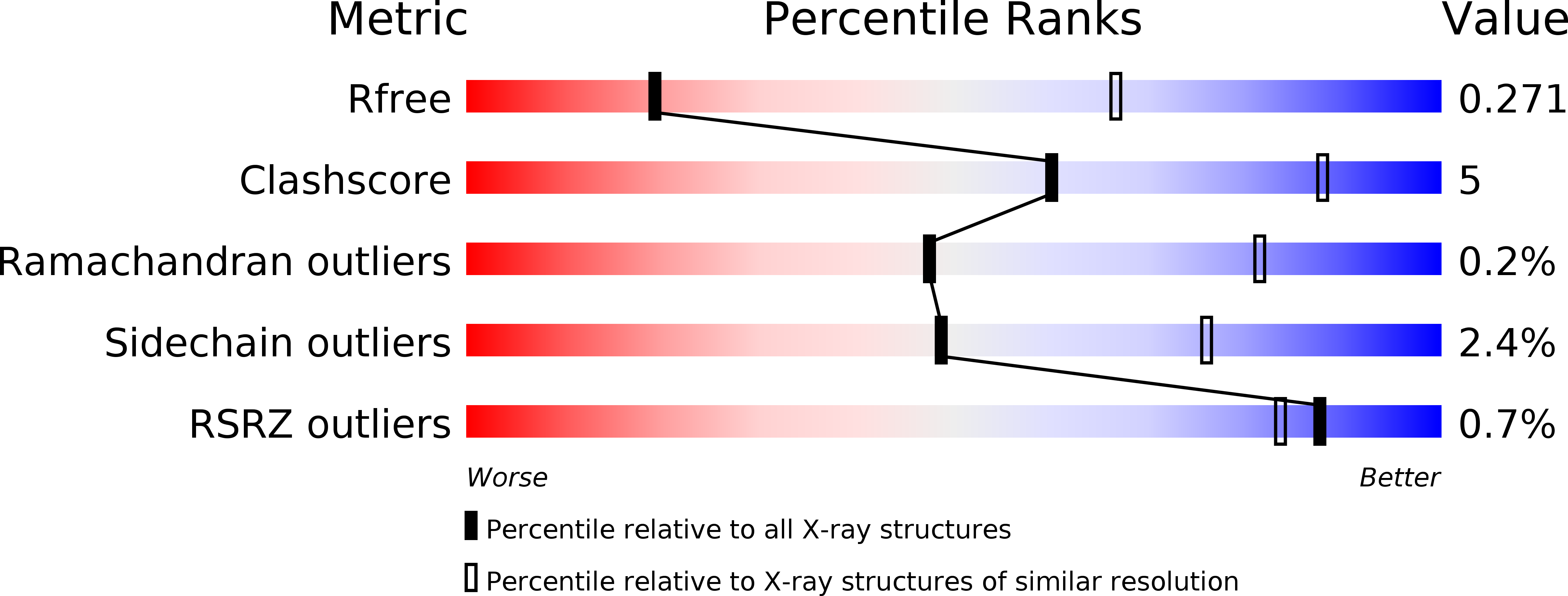
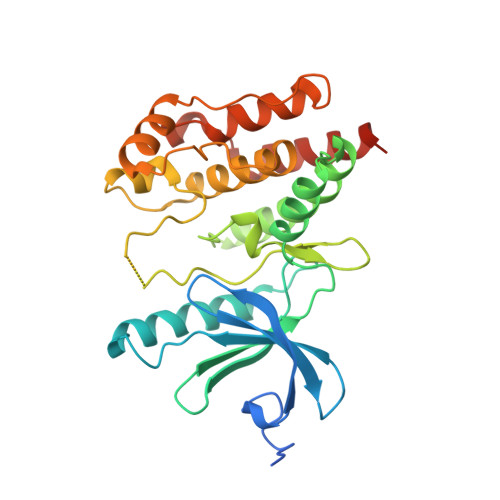
Nek7 is a serine/threonine-protein kinase required for proper spindle formation and cytokinesis. Elevated Nek7 levels have been observed in several cancers, and inhibition of Nek7 might provide a route to the development of cancer therapeutics. To date, no selective and potent Nek7 inhibitors have been identified. Nek7 crystal structures exhibit an improperly formed regulatory-spine (R-spine), characteristic of an inactive kinase. We reasoned that the preference of Nek7 to crystallise in this inactive conformation might hinder attempts to capture Nek7 in complex with Type I inhibitors. Here, we have introduced aromatic residues into the R-spine of Nek7 with the aim to stabilise the active conformation of the kinase through R-spine stacking. The strong R-spine mutant Nek7SRS retained catalytic activity and was crystallised in complex with compound 51, an ATP-competitive inhibitor of Nek2 and Nek7. Subsequently, we obtained the same crystal form for wild-type Nek7WT in apo form and bound to compound 51. The R-spines of the three well-ordered Nek7WT molecules exhibit variable conformations while the R-spines of the Nek7SRS molecules all have the same, partially stacked configuration. Compound 51 bound to Nek2 and Nek7 in similar modes, but differences in the precise orientation of a substituent highlights features that could be exploited in designing inhibitors that are selective for particular Nek family members. Although the SRS mutations are not required to obtain a Nek7-inhibitor structure, we conclude that it is a useful strategy for restraining the conformation of a kinase in order to promote crystallogenesis.
Organizational Affiliation:
Astbury Centre for Structural Molecular Biology, School of Molecular and Cellular Biology, Faculty of Biological Sciences, University of Leeds, Leeds, U.K.