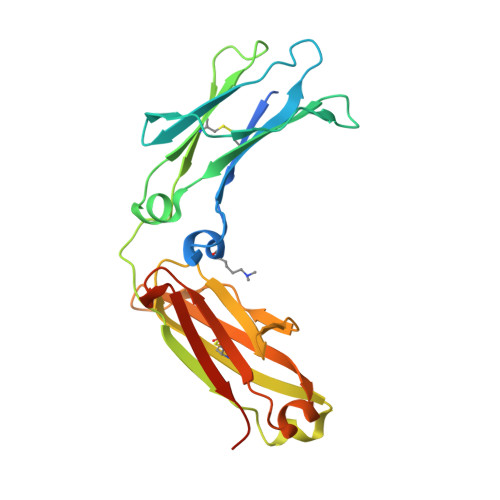
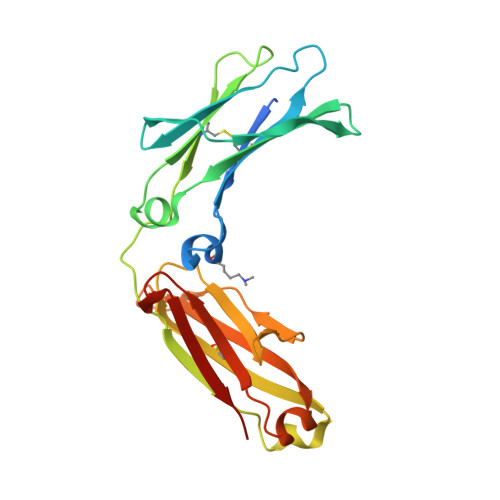
A single mutation increases heavy chain heterodimer assembly of bispecific antibodies by inducing structural disorder in one homodimer species.
Stutz, C., Blein, S.(2020) J Biological Chem
- PubMed: 32404368
- DOI: https://doi.org/10.1074/jbc.RA119.012335
- Primary Citation of Related Structures:
6G1E - PubMed Abstract:
We previously reported efficient heavy-chain assembly of heterodimeric bispecific antibodies by exchanging the interdomain protein interface of the human IgG1 CH3 dimer with the protein interface of the constant α and β domains of the human T-cell receptor, a technology known as bispecific engagement by antibodies based on the T-cell receptor (BEAT). Efficient heterodimerization in mammalian cell transient transfections was observed, but levels were influenced by the nature of the binding arms, particularly in the Fab-scFv-Fc format. In this study, we report a single amino acid change that significantly and consistently improved the heterodimerization rate of this format (≥95%) by inducing partial disorder in one homodimer species without affecting the heterodimer. Correct folding and assembly of the heterodimer were confirmed by the high-resolution (1.88-1.98 Å) crystal structure presented here. Thermal stability and 1-anilinonaphthalene-8-sulfonic acid-binding experiments, comparing original BEAT, mutated BEAT, and "knobs-into-holes" interfaces, suggested a cooperative assembly process of heavy chains in heterodimers. The observed gain in stability of the interfaces could be classified in the following rank order: mutated BEAT > original BEAT > knobs-into-holes. We therefore propose that the superior cooperativity found in BEAT interfaces is the key driver of their greater performance. Furthermore, we show how the mutated BEAT interface can be exploited for the routine preparation of drug candidates, with minimal risk of homodimer contamination using a single Protein A chromatography step.
- Department of Antibody Engineering, Ichnos Sciences S.A., Biopôle Lausanne-Epalinges, Epalinges, Switzerland.
Organizational Affiliation: