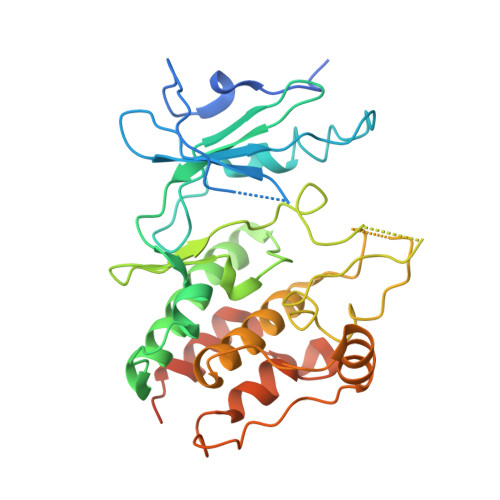
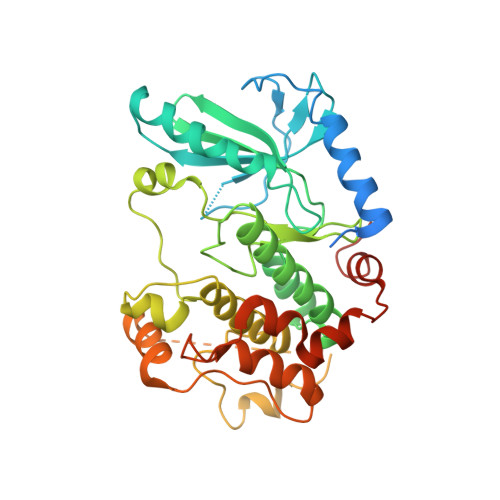
Small molecule stabilization of the KSR inactive state antagonizes oncogenic Ras signalling.
Dhawan, N.S., Scopton, A.P., Dar, A.C.(2016) Nature 537: 112-116
- PubMed: 27556948
- DOI: https://doi.org/10.1038/nature19327
- Primary Citation of Related Structures:
5KKR - PubMed Abstract:
Deregulation of the Ras-mitogen activated protein kinase (MAPK) pathway is an early event in many different cancers and a key driver of resistance to targeted therapies. Sustained signalling through this pathway is caused most often by mutations in K-Ras, which biochemically favours the stabilization of active RAF signalling complexes. Kinase suppressor of Ras (KSR) is a MAPK scaffold that is subject to allosteric regulation through dimerization with RAF. Direct targeting of KSR could have important therapeutic implications for cancer; however, testing this hypothesis has been difficult owing to a lack of small-molecule antagonists of KSR function. Guided by KSR mutations that selectively suppress oncogenic, but not wild-type, Ras signalling, we developed a class of compounds that stabilize a previously unrecognized inactive state of KSR. These compounds, exemplified by APS-2-79, modulate KSR-dependent MAPK signalling by antagonizing RAF heterodimerization as well as the conformational changes required for phosphorylation and activation of KSR-bound MEK (mitogen-activated protein kinase kinase). Furthermore, APS-2-79 increased the potency of several MEK inhibitors specifically within Ras-mutant cell lines by antagonizing release of negative feedback signalling, demonstrating the potential of targeting KSR to improve the efficacy of current MAPK inhibitors. These results reveal conformational switching in KSR as a druggable regulator of oncogenic Ras, and further suggest co-targeting of enzymatic and scaffolding activities within Ras-MAPK signalling complexes as a therapeutic strategy for overcoming Ras-driven cancers.
Organizational Affiliation:
Department of Oncological Sciences, The Tisch Cancer Institute, The Icahn School of Medicine at Mount Sinai, New York, New York 10029, USA.