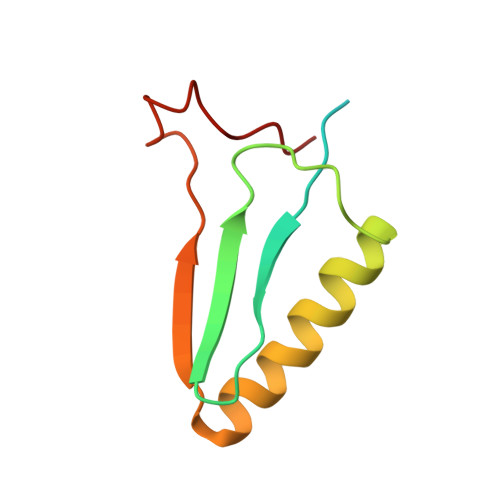
The NMR solution structure and function of RPA3313: a putative ribosomal transport protein from Rhodopseudomonas palustris.
Catazaro, J., Lowe, A.J., Cerny, R.L., Powers, R.(2017) Proteins 85: 93-102
- PubMed: 27802574
- DOI: https://doi.org/10.1002/prot.25201
- Primary Citation of Related Structures:
5JN6 - PubMed Abstract:
Protein function elucidation often relies heavily on amino acid sequence analysis and other bioinformatics approaches. The reliance is extended to structure homology modeling for ligand docking and protein-protein interaction mapping. However, sequence analysis of RPA3313 exposes a large, unannotated class of hypothetical proteins mostly from the Rhizobiales order. In the absence of sequence and structure information, further functional elucidation of this class of proteins has been significantly hindered. A high quality NMR structure of RPA3313 reveals that the protein forms a novel split ββαβ fold with a conserved ligand binding pocket between the first β-strand and the N-terminus of the α-helix. Conserved residue analysis and protein-protein interaction prediction analyses reveal multiple protein binding sites and conserved functional residues. Results of a mass spectrometry proteomic analysis strongly point toward interaction with the ribosome and its subunits. The combined structural and proteomic analyses suggest that RPA3313 by itself or in a larger complex may assist in the transportation of substrates to or from the ribosome for further processing. Proteins 2016; 85:93-102. © 2016 Wiley Periodicals, Inc.
Organizational Affiliation:
Department of Chemistry, University of Nebraska-Lincoln, Lincoln, Nebraska, 68588-0304.