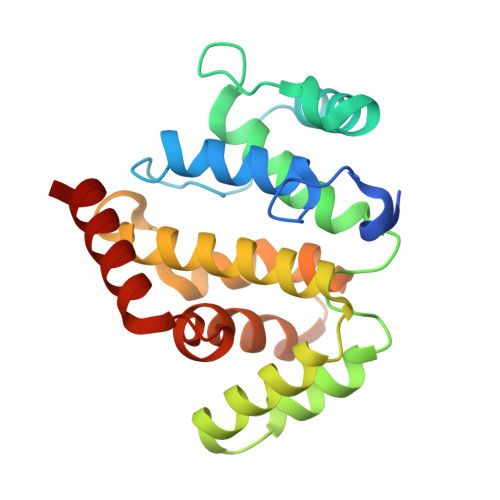
Structural Basis for the Specific Recognition of RhoA by the Dual GTPase-activating Protein ARAP3
Bao, H., Li, F., Wang, C., Wang, N., Jiang, Y., Tang, Y., Wu, J., Shi, Y.(2016) J Biol Chem 291: 16709-16719
- PubMed: 27311713
- DOI: https://doi.org/10.1074/jbc.M116.736140
- Primary Citation of Related Structures:
5JCP, 5JD0 - PubMed Abstract:
ARAP3 (Arf-GAP with Rho-GAP domain, ANK repeat, and PH domain-containing protein 3) is unique for its dual specificity GAPs (GTPase-activating protein) activity for Arf6 (ADP-ribosylation factor 6) and RhoA (Ras homolog gene family member A) regulated by phosphatidylinositol 3,4,5-trisphosphate and a small GTPase Rap1-GTP and is involved in regulation of cell shape and adhesion. However, the molecular interface between the ARAP3-RhoGAP domain and RhoA is unknown, as is the substrates specificity of the RhoGAP domain. In this study, we solved the crystal structure of RhoA in complex with the RhoGAP domain of ARAP3. The structure of the complex presented a clear interface between the RhoGAP domain and RhoA. By analyzing the crystal structure and in combination with in vitro GTPase activity assays and isothermal titration calorimetry experiments, we identified the crucial residues affecting RhoGAP activity and substrates specificity among RhoA, Rac1 (Ras-related C3 botulinum toxin substrate 1), and Cdc42 (cell division control protein 42 homolog).
Organizational Affiliation:
From the Hefei National Laboratory for Physical Science at Microscale, Collaborative Innovation Center of Chemistry for Life Sciences and School of Life Sciences, University of Science and Technology of China, Hefei, Anhui, 230027, China.