An allosteric PRC2 inhibitor targeting the H3K27me3 binding pocket of
Qi, W., Zhao, K., Gu, J., Huang, Y., Wang, Y., Zhang, H., Zhang, M., Zhang, J., Yu, Z., Li, L., Teng, L., Chuai, S., Zhang, C., Zhao, M., Chan, H., Chen, Z., Fang, D., Fei, Q., Feng, L., Feng, L., Gao, Y., Ge, H., Ge, X., Li, G., Lingel, A., Lin, Y., Liu, Y., Luo, F., Shi, M., Wang, L., Wang, Z., Yu, Y., Zeng, J., Zeng, C., Zhang, L., Zhang, Q., Zhou, S., Oyang, C., Atadja, P., Li, E.(2017) Nat Chem Biol 13: 381-388
- PubMed: 28135235
- DOI: https://doi.org/10.1038/nchembio.2304
- Primary Citation of Related Structures:
5GSA - PubMed Abstract:
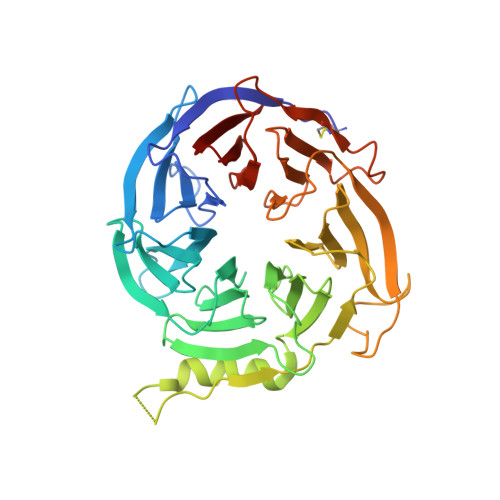
Polycomb repressive complex 2 (PRC2) consists of three core subunits, EZH2, EED and SUZ12, and plays pivotal roles in transcriptional regulation. The catalytic subunit EZH2 methylates histone H3 lysine 27 (H3K27), and its activity is further enhanced by the binding of EED to trimethylated H3K27 (H3K27me3). Small-molecule inhibitors that compete with the cofactor S-adenosylmethionine (SAM) have been reported. Here we report the discovery of EED226, a potent and selective PRC2 inhibitor that directly binds to the H3K27me3 binding pocket of EED. EED226 induces a conformational change upon binding EED, leading to loss of PRC2 activity. EED226 shows similar activity to SAM-competitive inhibitors in blocking H3K27 methylation of PRC2 target genes and inducing regression of human lymphoma xenograft tumors. Interestingly, EED226 also effectively inhibits PRC2 containing a mutant EZH2 protein resistant to SAM-competitive inhibitors. Together, we show that EED226 inhibits PRC2 activity via an allosteric mechanism and offers an opportunity for treatment of PRC2-dependent cancers.
Organizational Affiliation:
Novartis Institutes for BioMedical Research, Shanghai, China.