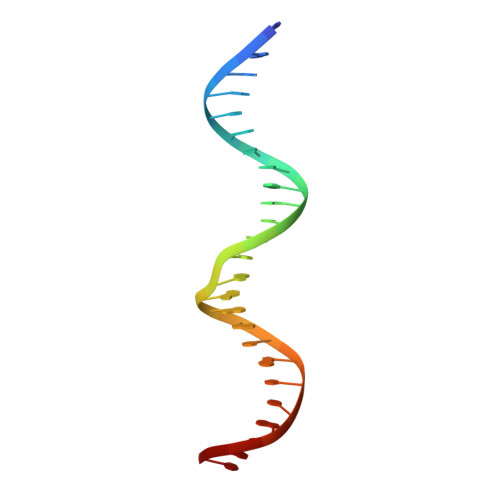
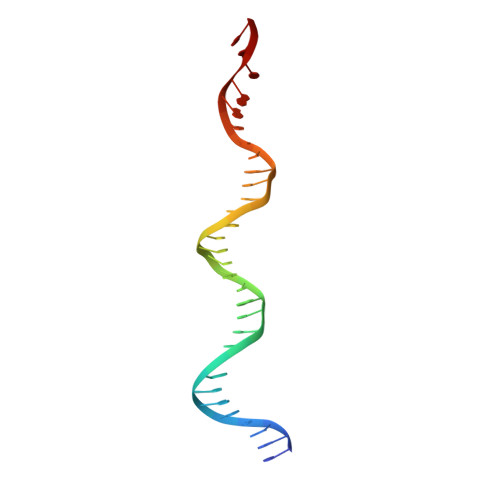
Structural role of the flanking DNA in mariner transposon excision.
Dornan, J., Grey, H., Richardson, J.M.(2015) Nucleic Acids Res 43: 2424-2432
- PubMed: 25662605
- DOI: https://doi.org/10.1093/nar/gkv096
- Primary Citation of Related Structures:
4U7B - PubMed Abstract:
During cut-and-paste mariner/Tc1 transposition, transposon DNA is cut precisely at its junction with flanking DNA, ensuring the transposon is neither shortened nor lengthened with each transposition event. Each transposon end is flanked by a TpA dinucleotide: the signature target site duplication of mariner/Tc1 transposition. To establish the role of this sequence in accurate DNA cleavage, we have determined the crystal structure of a pre-second strand cleavage mariner Mos1 transpososome. The structure reveals the route of an intact DNA strand through the transposase active site before second strand cleavage. The crossed architecture of this pre-second strand cleavage paired-end complex supports our proposal that second strand cleavage occurs in trans. The conserved mariner transposase WVPHEL and YSPDL motifs position the strand for accurate DNA cleavage. Base-specific recognition of the flanking DNA by conserved amino acids is revealed, defining a new role for the WVPHEL motif in mariner transposition and providing a molecular explanation for in vitro mutagenesis data. Comparison of the pre-TS cleavage and post-cleavage Mos1 transpososomes with structures of Prototype Foamy Virus intasomes suggests a binding mode for target DNA prior to Mos1 transposon integration.
Organizational Affiliation:
Institute of Structural and Molecular Biology, School of Biological Sciences, University of Edinburgh, The King's Buildings, Max Born Crescent, Edinburgh EH9 3BF, UK.