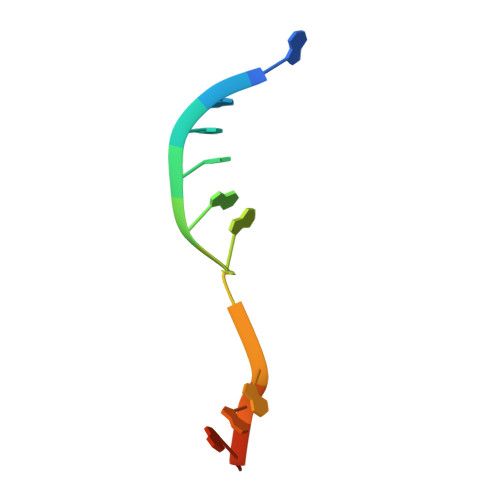
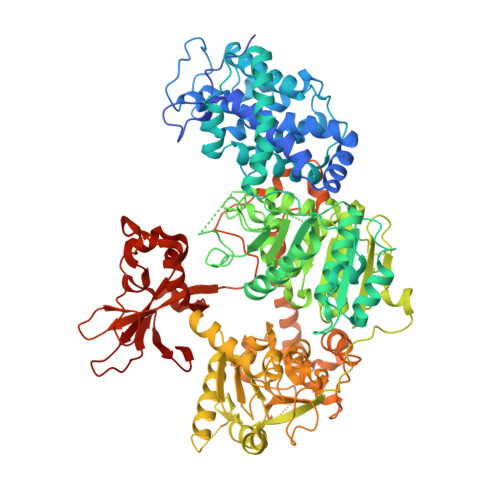
Structures of CRISPR Cas3 offer mechanistic insights into Cascade-activated DNA unwinding and degradation.
Huo, Y., Nam, K.H., Ding, F., Lee, H., Wu, L., Xiao, Y., Farchione, M.D., Zhou, S., Rajashankar, K., Kurinov, I., Zhang, R., Ke, A.(2014) Nat Struct Mol Biol 21: 771-777
- PubMed: 25132177
- DOI: https://doi.org/10.1038/nsmb.2875
- Primary Citation of Related Structures:
4QQW, 4QQX, 4QQY, 4QQZ - PubMed Abstract:
CRISPR drives prokaryotic adaptation to invasive nucleic acids such as phages and plasmids, using an RNA-mediated interference mechanism. Interference in type I CRISPR-Cas systems requires a targeting Cascade complex and a degradation machine, Cas3, which contains both nuclease and helicase activities. Here we report the crystal structures of Thermobifida fusca Cas3 bound to single-stranded (ss) DNA substrate and show that it is an obligate 3'-to-5' ssDNase that preferentially accepts substrate directly from the helicase moiety. Conserved residues in the HD-type nuclease coordinate two irons for ssDNA cleavage. We demonstrate ATP coordination and conformational flexibility of the SF2-type helicase domain. Cas3 is specifically guided toward Cascade-bound target DNA by a PAM sequence, through physical interactions with both the nontarget substrate strand and the CasA protein. The sequence of recognition events ensures well-controlled DNA targeting and degradation of foreign DNA by Cascade and Cas3.
- 1] Department of Molecular Biology and Genetics, Cornell University, Ithaca, New York, USA. [2].
Organizational Affiliation: