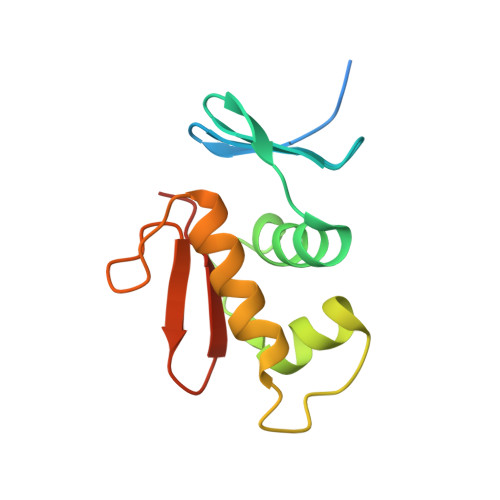
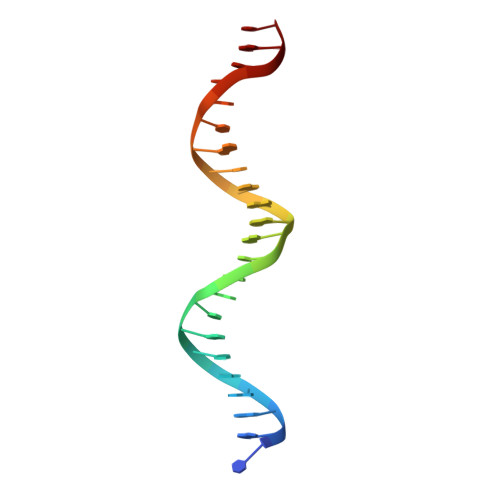
Structural dynamics of the two-component response regulator RstA in recognition of promoter DNA element.
Li, Y.C., Chang, C.K., Chang, C.F., Cheng, Y.H., Fang, P.J., Yu, T., Chen, S.C., Li, Y.C., Hsiao, C.D., Huang, T.H.(2014) Nucleic Acids Res 42: 8777-8788
- PubMed: 24990372
- DOI: https://doi.org/10.1093/nar/gku572
- Primary Citation of Related Structures:
2MLK, 4NHJ, 4NIC - PubMed Abstract:
The RstA/RstB system is a bacterial two-component regulatory system consisting of the membrane sensor, RstB and its cognate response regulator (RR) RstA. The RstA of Klebsiella pneumoniae (kpRstA) consists of an N-terminal receiver domain (RD, residues 1-119) and a C-terminal DNA-binding domain (DBD, residues 130-236). Phosphorylation of kpRstA induces dimerization, which allows two kpRstA DBDs to bind to a tandem repeat, called the RstA box, and regulate the expression of downstream genes. Here we report the solution and crystal structures of the free kpRstA RD, DBD and DBD/RstA box DNA complex. The structure of the kpRstA DBD/RstA box complex suggests that the two protomers interact with the RstA box in an asymmetric fashion. Equilibrium binding studies further reveal that the two protomers within the kpRstA dimer bind to the RstA box in a sequential manner. Taken together, our results suggest a binding model where dimerization of the kpRstA RDs provides the platform to allow the first kpRstA DBD protomer to anchor protein-DNA interaction, whereas the second protomer plays a key role in ensuring correct recognition of the RstA box.
Organizational Affiliation:
Institute of Molecular Biology, Academia Sinica, Taipei 115, Taiwan, ROC Institute of Bioinformatics and Structural Biology, National Tsing Hua University, Hsinchu 300, Taiwan.