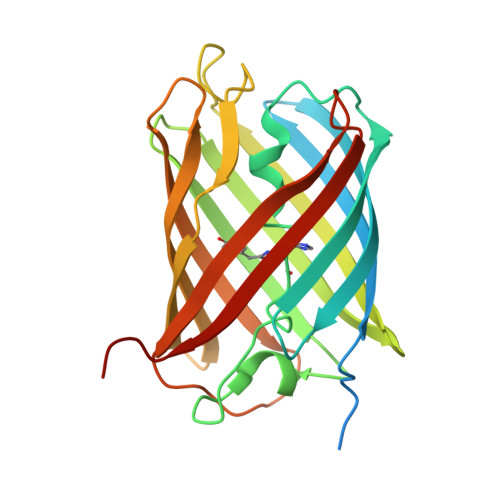
Green-to-red photoconvertible Dronpa mutant for multimodal super-resolution fluorescence microscopy
Moeyaert, B., Nguyen Bich, N., De Zitter, E., Rocha, S., Clays, K., Mizuno, H., van Meervelt, L., Hofkens, J., Dedecker, P.(2014) ACS Nano 8: 1664-1673
- PubMed: 24410188
- DOI: https://doi.org/10.1021/nn4060144
- Primary Citation of Related Structures:
4HQ8, 4HQ9, 4HQC, 4IZN - PubMed Abstract:
Advanced imaging techniques crucially depend on the labels used. In this work, we present the structure-guided design of a fluorescent protein that displays both reversibly photochromic and green-to-red photoconversion behavior. We first designed ffDronpa, a mutant of the photochromic fluorescent protein Dronpa that matures up to three times faster while retaining its interesting photochromic features. Using a combined evolutionary and structure-driven rational design strategy, we developed a green-to-red photoconvertible ffDronpa mutant, called pcDronpa, and explored different optimization strategies that resulted in its improved version, pcDronpa2. This fluorescent probe combines a high brightness with low photobleaching and photoblinking. We herein show that, despite its tetrameric nature, pcDronpa2 allows for multimodal subdiffraction imaging by sequentially imaging a given sample using both super-resolution fluctuation imaging and localization microscopy.
- Department of Chemistry, KU Leuven , Celestijnenlaan 200F, bus 2404, 3001 Heverlee, Belgium.
Organizational Affiliation: