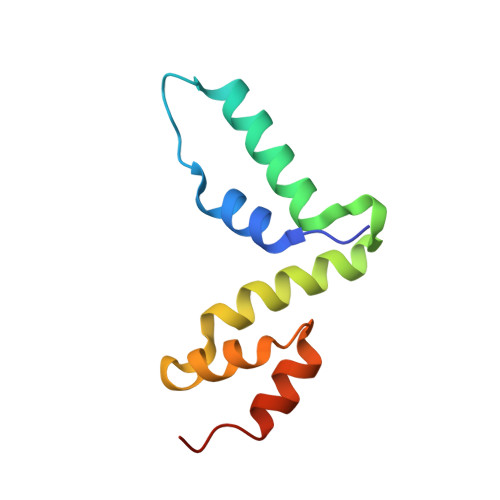
Structure of the Scan Domain of Human Paternally Expressed Gene 3 Protein.
Rimsa, V., Eadsforth, T.C., Hunter, W.N.(2013) PLoS One 8: 69538
- PubMed: 23936039
- DOI: https://doi.org/10.1371/journal.pone.0069538
- Primary Citation of Related Structures:
4BHX - PubMed Abstract:
Human paternally expressed gene 3 protein (PEG3) is a large multi-domain entity with diverse biological functions, including acting as a transcription factor. PEG3 contains twelve Cys2-His2 type zinc finger domains, extended regions of predicted disorder and at the N-terminus a SCAN domain. PEG3 has been identified as partner of the E3 ubiquitin-protein ligase Siah1, an association we sought to investigate. An efficient bacterial recombinant expression system of the human PEG3-SCAN domain was prepared and crystals appeared spontaneously when the protein was being concentrated after purification. The structure was determined at 1.95 Å resolution and reveals a polypeptide fold of five helices in an extended configuration. An extensive dimerization interface, using almost a quarter of the solvent accessible surface, and key salt bridge interactions explain the stability of the dimer. Comparison with other SCAN domains reveals a high degree of conservation involving residues that contribute to the dimer interface. The PEG3-SCAN domain appears to constitute an assembly block, enabling PEG3 homo- or heterodimerization to control gene expression in a combinatorial fashion.
Organizational Affiliation:
Division of Biological Chemistry and Drug Discovery, College of Life Sciences, University of Dundee, Dundee, United Kingdom.