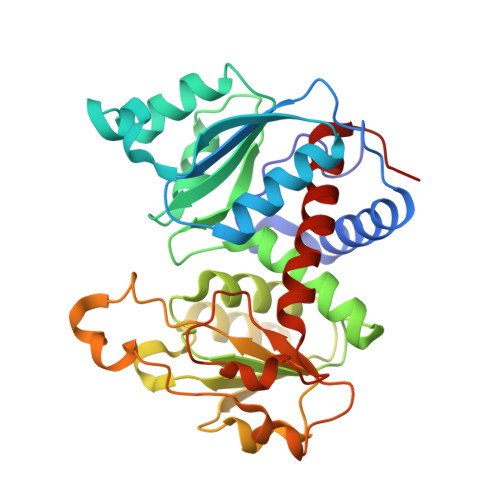
A second allosteric site in Escherichia coli aspartate transcarbamoylase.
Peterson, A.W., Cockrell, G.M., Kantrowitz, E.R.(2012) Biochemistry 51: 4776-4778
- PubMed: 22667327
- DOI: https://doi.org/10.1021/bi3006219
- Primary Citation of Related Structures:
4F04 - PubMed Abstract:
Escherichia coli aspartate transcarbamoylase is feedback inhibited by CTP and UTP in the presence of CTP. Here, we show by X-ray crystallography that UTP binds to a unique site on each regulatory chain of the enzyme that is near but not overlapping with the known CTP site. These results bring into question all of the previously proposed mechanisms of allosteric regulation in aspartate transcarbamoylase.
- Department of Chemistry, Merkert Chemistry Center, Boston College, Chestnut Hill, Massachusetts 02467, USA.
Organizational Affiliation: