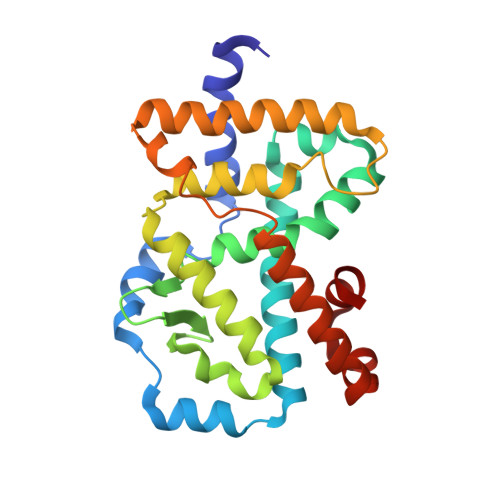
Structural basis for hydroxycholesterols as natural ligands of orphan nuclear receptor RORgamma.
Jin, L., Martynowski, D., Zheng, S., Wada, T., Xie, W., Li, Y.(2010) Mol Endocrinol 24: 923-929
- PubMed: 20203100
- DOI: https://doi.org/10.1210/me.2009-0507
- Primary Citation of Related Structures:
3KYT, 3L0J, 3L0L - PubMed Abstract:
The retinoic acid-related orphan receptor gamma (RORgamma) has important roles in development and metabolic homeostasis. Although the biological functions of RORgamma have been studied extensively, no ligands for RORgamma have been identified, and no structure of RORgamma has been reported. In this study, we showed that hydroxycholesterols promote the recruitment of coactivators by RORgamma using biochemical assays. We also report the crystal structures of the RORgamma ligand-binding domain bound with hydroxycholesterols. The structures reveal the binding modes of various hydroxycholesterols in the RORgamma pocket, with the receptors all adopting the canonical active conformation. Mutations that disrupt the binding of hydroxycholesterols abolish the constitutive activity of RORgamma. Our observations suggest an important role for the endogenous hydroxycholesterols in modulating RORgamma-dependent biological processes.
Organizational Affiliation:
Key Laboratory for Cell Biology and Tumor Cell Engineering of the Ministry of Education, Xiamen University, Fujian 361005, China