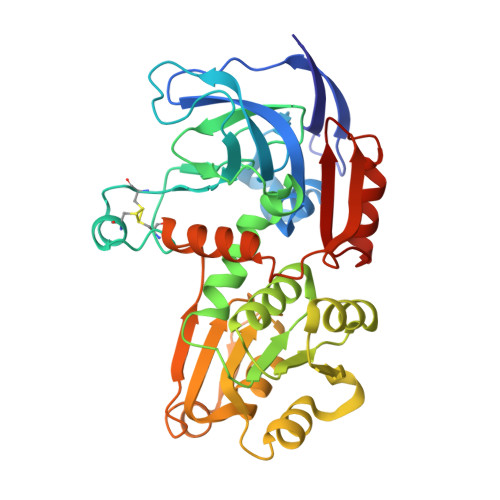
Structure and function of the l-threonine dehydrogenase (TkTDH) from the hyperthermophilic archaeon Thermococcus kodakaraensis
Bowyer, A., Mikolajek, H., Stuart, J.W., Wood, S.P., Jamil, F., Rashid, N., Akhtar, M., Cooper, J.B.(2009) J Struct Biol 168: 294-304
- PubMed: 19616102
- DOI: https://doi.org/10.1016/j.jsb.2009.07.011
- Primary Citation of Related Structures:
3GFB - PubMed Abstract:
The X-ray structure of the holo-form of l-threonine dehydrogenase (TDH) from Thermococcus kodakaraensis (TkTDH) has been determined at 2.4A resolution. TDH catalyses the NAD(+)-dependent oxidation of l-threonine to 2-amino-3-ketobutyrate, and is one of the first enzymes in this family to be solved by X-ray crystallography. The enzyme is a homo-tetramer, each monomer consisting of 350 amino acids that form two domains; a catalytic domain and a nicotinamide co-factor (NAD(+))-binding domain, which contains an alpha/beta Rossmann fold motif. An extended twelve-stranded beta-sheet is formed by the association of pairs of monomers in the tetramer. TkTDH shows strong overall structural similarity to TDHs from thermophiles and alcohol dehydrogenases (ADH) from lower life forms, despite low sequence homology, exhibiting the same overall fold of the monomer and assembly of the tetramer. The structure reveals the binding site of the essential co-factor NAD(+) which is present in all subunits. Docking studies suggest a mode of interaction of TDH with 2-amino-3-ketobutyrate CoA ligase, the subsequent enzyme in the pathway for conversion of threonine to glycine. TDH is known to form a stable functional complex with 2-amino-3-ketobutyrate ligase, most probably to shield an unstable intermediate.
Organizational Affiliation:
University of Southampton, UK.