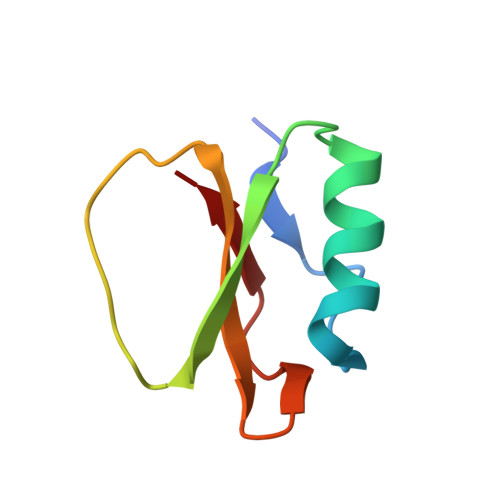
Refinement of the three-dimensional solution structure of barley serine proteinase inhibitor 2 and comparison with the structures in crystals.
Ludvigsen, S., Shen, H.Y., Kjaer, M., Madsen, J.C., Poulsen, F.M.(1991) J Mol Biol 222: 621-635
- PubMed: 1748996
- DOI: https://doi.org/10.1016/0022-2836(91)90500-6
- Primary Citation of Related Structures:
3CI2 - PubMed Abstract:
The three-dimensional structure of barley serine proteinase inhibitor, CI-2, has been determined using nuclear magnetic resonance spectroscopy. The present structure determination is a refinement of the structure previously determined by us, using in the present case stereo-specific assignments, and a virtually complete set of assignments of the two-dimensional nuclear Overhauser spectrum. The structure determination is based on the identification of more than 1300 nuclear Overhauser effects, of which 961 were used in the structure calculation as distance restraints, and on 94 dihedral angle restraints, of which 31 are for chi 1 angles in defined chiral centers. These have been used to calculate a series of 20 three-dimensional structures using a combination of distance geometry, simulated annealing and restrained molecular dynamics. Each of the 20 structures was in agreement within less than 0.5 A of each of the distance restraints and with all dihedral angle restraints. When compared to the geometric average structure of the 20 refined structures the root-mean-square differences for the backbone atoms were 0.8 (+/- 0.2) A and for all atoms were 1.6 (+/- 0.2) A. By comparison, the values obtained for the structures determined previously were 1.4 (+/- 0.2) A and 2.1 (+/- 0.1) A, respectively. The structures were also compared to the structure determined in the crystalline state by X-ray diffraction showing root-mean-square differences of 1.6 (+/- 0.2) A and 2.8 (+/- 0.2) A for the backbone and all atoms, respectively. Common features of the solution structure and the two crystal structures are the four-stranded beta-structure, composed of a pair of parallel strands, and three pairs of antiparallel beta-strands flanked on one side by a 12-residue alpha-helix and on the other side by a loop containing the serine proteinase binding site. The new analysis of the structure has revealed an additional pair of antiparallel beta-strands, consisting of residues 65 to 67 and 81 to 83, that was not seen in either of the crystal structures or the previous solution structure. Identification of this was based on nuclear magnetic resonance evidence for the hydrogen bond (67HN to 81CO) not reported previously. Also the presence of a bifurcated hydrogen bond involving Phe69 CO and HN atoms of Ala77 and Gln78 was observed in solution but not in crystals. Minor differences between the two structures were observed in the phi-angles of residues Met59 and Glu60 in the inhibitory site.
Organizational Affiliation:
Kemisk Afdeling, Carlsberg Laboratorium, Valby Copenhagen, Denmark.