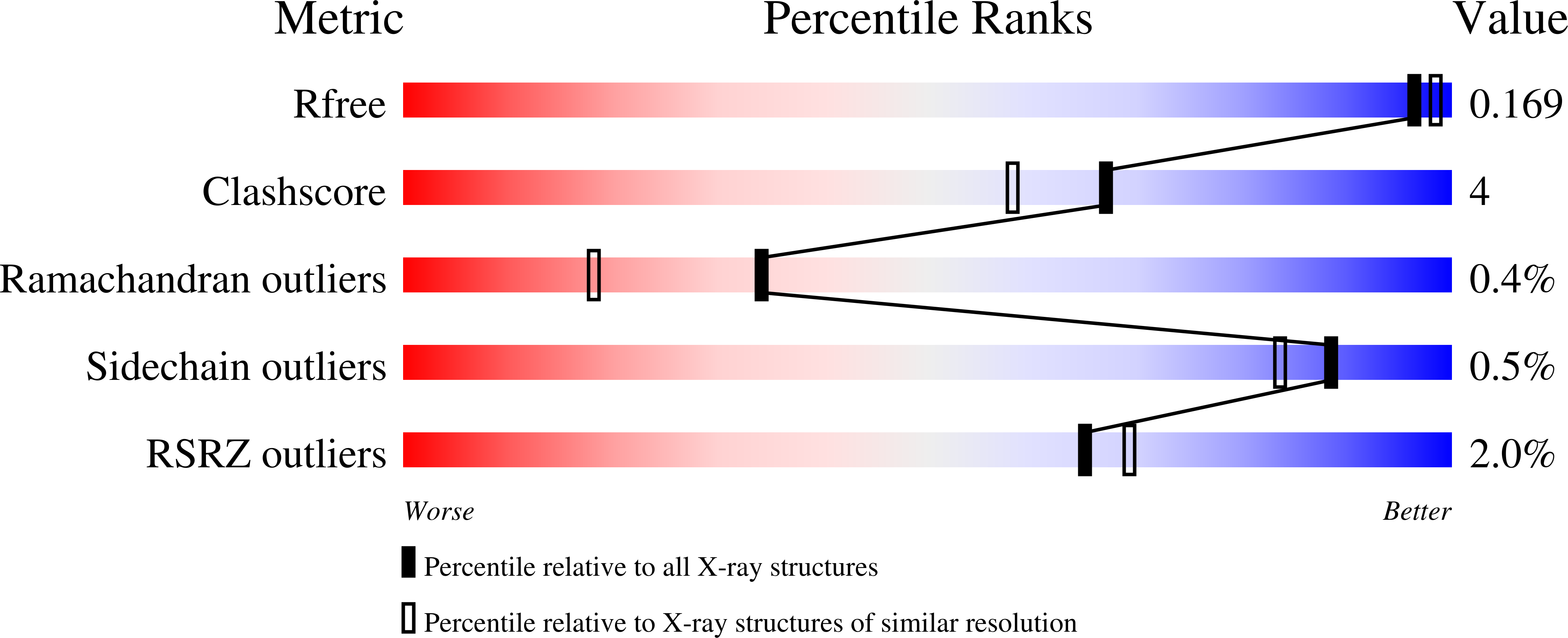
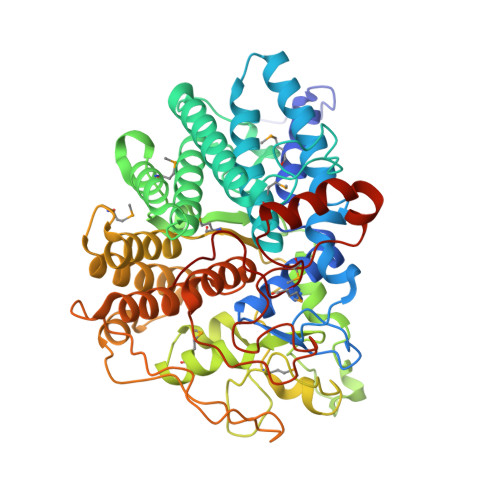
Structure of BT_3984, a member of the SusD/RagB family of nutrient-binding molecules.
Bakolitsa, C., Xu, Q., Rife, C.L., Abdubek, P., Astakhova, T., Axelrod, H.L., Carlton, D., Chen, C., Chiu, H.J., Clayton, T., Das, D., Deller, M.C., Duan, L., Ellrott, K., Farr, C.L., Feuerhelm, J., Grant, J.C., Grzechnik, A., Han, G.W., Jaroszewski, L., Jin, K.K., Klock, H.E., Knuth, M.W., Kozbial, P., Krishna, S.S., Kumar, A., Lam, W.W., Marciano, D., McMullan, D., Miller, M.D., Morse, A.T., Nigoghossian, E., Nopakun, A., Okach, L., Puckett, C., Reyes, R., Tien, H.J., Trame, C.B., van den Bedem, H., Weekes, D., Hodgson, K.O., Wooley, J., Elsliger, M.A., Deacon, A.M., Godzik, A., Lesley, S.A., Wilson, I.A.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 1274-1280
- PubMed: 20944222
- DOI: https://doi.org/10.1107/S1744309110032999
- Primary Citation of Related Structures:
3CGH - PubMed Abstract:
The crystal structure of the Bacteroides thetaiotaomicron protein BT_3984 was determined to a resolution of 1.7 Å and was the first structure to be determined from the extensive SusD family of polysaccharide-binding proteins. SusD is an essential component of the sus operon that defines the paradigm for glycan utilization in dominant members of the human gut microbiota. Structural analysis of BT_3984 revealed an N-terminal region containing several tetratricopeptide repeats (TPRs), while the signature C-terminal region is less structured and contains extensive loop regions. Sequence and structure analysis of BT_3984 suggests the presence of binding interfaces for other proteins from the polysaccharide-utilization complex.
Organizational Affiliation:
Program on Bioinformatics and Systems Biology, Sanford-Burnham Medical Research Institute, La Jolla, CA, USA.