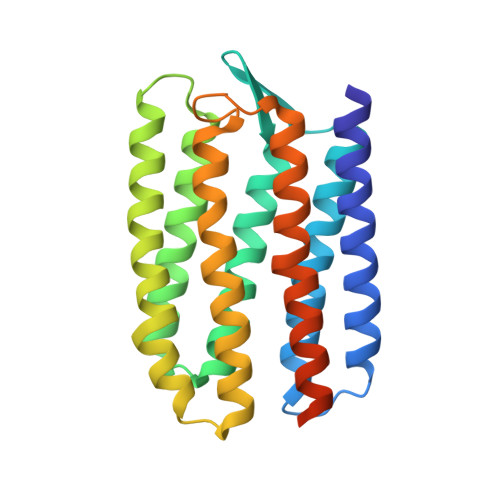
Active State of Sensory Rhodopsin II: Structural Determinants for Signal Transfer and Proton Pumping.
Gushchin, I., Reshetnyak, A., Borshchevskiy, V., Ishchenko, A., Round, E., Grudinin, S., Engelhard, M., Buldt, G., Gordeliy, V.(2011) J Mol Biology 412: 591-600
- PubMed: 21840321
- DOI: https://doi.org/10.1016/j.jmb.2011.07.022
- Primary Citation of Related Structures:
3QAP, 3QDC - PubMed Abstract:
The molecular mechanism of transmembrane signal transduction is still a pertinent question in cellular biology. Generally, a receptor can transfer an external signal via its cytoplasmic surface, as found for G-protein-coupled receptors such as rhodopsin, or via the membrane domain, such as that in sensory rhodopsin II (SRII) in complex with its transducer, HtrII. In the absence of HtrII, SRII functions as a proton pump. Here, we report on the crystal structure of the active state of uncomplexed SRII from Natronomonas pharaonis, NpSRII. The problem with a dramatic loss of diffraction quality upon loading of the active state was overcome by growing better crystals and by reducing the occupancy of the state. The conformational changes in the region comprising helices F and G are similar to those observed for the NpSRII-transducer complex but are much more pronounced. The meaning of these differences for the understanding of proton pumping and signal transduction by NpSRII is discussed.
- Laboratoire des Protéines Membranaires, Institut de Biologie Structurale J.-P. Ebel, UMR5075 CEA-CNRS-UJF, 38027 Grenoble, France.
Organizational Affiliation: