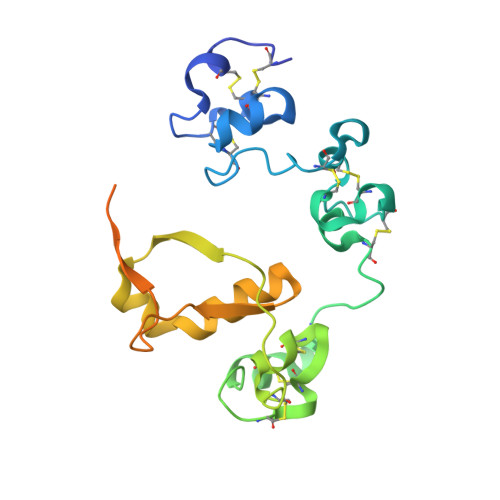
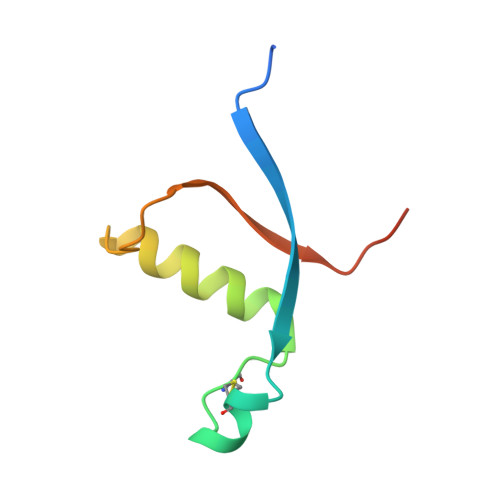
Effects of S1 cleavage on the structure, surface export, and signaling activity of human Notch1 and Notch2.
Gordon, W.R., Vardar-Ulu, D., L'Heureux, S., Ashworth, T., Malecki, M.J., Sanchez-Irizarry, C., McArthur, D.G., Histen, G., Mitchell, J.L., Aster, J.C., Blacklow, S.C.(2009) PLoS One 4: e6613-e6613
- PubMed: 19701457
- DOI: https://doi.org/10.1371/journal.pone.0006613
- Primary Citation of Related Structures:
3I08 - PubMed Abstract:
Notch receptors are normally cleaved during maturation by a furin-like protease at an extracellular site termed S1, creating a heterodimer of non-covalently associated subunits. The S1 site lies within a key negative regulatory region (NRR) of the receptor, which contains three highly conserved Lin12/Notch repeats and a heterodimerization domain (HD) that interact to prevent premature signaling in the absence of ligands. Because the role of S1 cleavage in Notch signaling remains unresolved, we investigated the effect of S1 cleavage on the structure, surface trafficking and ligand-mediated activation of human Notch1 and Notch2, as well as on ligand-independent activation of Notch1 by mutations found in human leukemia. The X-ray structure of the Notch1 NRR after furin cleavage shows little change when compared with that of an engineered Notch1 NRR lacking the S1-cleavage loop. Likewise, NMR studies of the Notch2 HD domain show that the loop containing the S1 site can be removed or cleaved without causing a substantial change in its structure. However, Notch1 and Notch2 receptors engineered to resist S1 cleavage exhibit unexpected differences in surface delivery and signaling competence: S1-resistant Notch1 receptors exhibit decreased, but detectable, surface expression and ligand-mediated receptor activation, whereas S1-resistant Notch2 receptors are fully competent for cell surface delivery and for activation by ligands. Variable dependence on S1 cleavage also extends to T-ALL-associated NRR mutations, as common class 1 mutations display variable decrements in ligand-independent activation when introduced into furin-resistant receptors, whereas a class 2 mutation exhibits increased signaling activity. S1 cleavage has distinct effects on the surface expression of Notch1 and Notch2, but is not generally required for physiologic or pathophysiologic activation of Notch proteins. These findings are consistent with models for receptor activation in which ligand-binding or T-ALL-associated mutations lead to conformational changes of the NRR that permit metalloprotease cleavage.
Organizational Affiliation:
Department of Pathology, Brigham and Women's Hospital and Harvard Medical School, Boston, Massachusetts, United States of America.