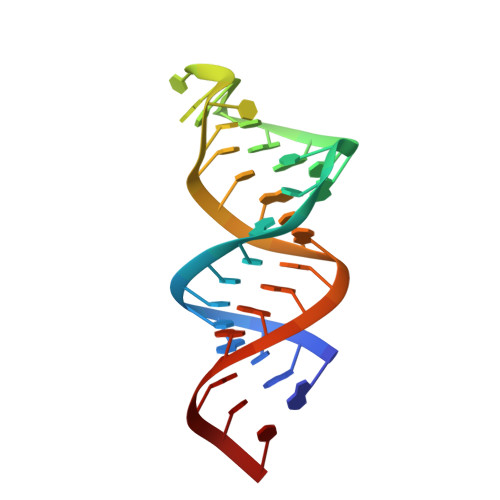
Solution structure of a reverse transcriptase recognition site of a LINE RNA from zebrafish.
Otsu, M., Kajikawa, M., Okada, N., Kawai, G.(2017) J Biochem 162: 279-285
- PubMed: 28431120
- DOI: https://doi.org/10.1093/jb/mvx026
- Primary Citation of Related Structures:
2RVO - PubMed Abstract:
Long interspersed nuclear element (LINE) is known to be transposed by reverse transcription using its RNA transcript. Recognition of the 3' stem-loop of LINE RNA by its reverse transcriptase (RT) is an important step of the retrotransposition. Our previous study revealed that the second G residue (G8) in the GGAUA loop of a 17mer LINE RNA from eel, UnaL2-17, is recognized by its RT and the U residue (U10) in the same loop is required to maintain the loop structure (Baba S, Kajikawa M, Okada N, Kawai G. Solution structure of an RNA stem-loop derived from the 3' conserved region of eel LINE UnaL2. RNA 2004;10:1380-1387). ZfL2-2, a LINE from zebrafish, has the same 3' stem-loop with UnaL2 and ZfL2-1 has similar but distinct 3' stem-loop with an insertion which can form an additional stem-loop. Here, we determined the solution structure of the 34mer RT recognition site of the LINE RNA (ZfL2-1-34). It was found that ZfL2-1-34 forms a hairpin with an internal loop, the tertiary structure of which is superimposed with that of ZfL2-2. It is noted that A10 and the inserted stem-loop, starting with A12, in ZfL2-1-34 located at the positions corresponding to those of G8 and U10, respectively, in UnaL2-17. These results strongly suggest that the two LINEs share the similar recognition mechanism and the A10 in ZfL2-1-34 is the determinant recognized by its RT.
- Department of Life and Environmental Sciences, Faculty of Engineering, Chiba Institute of Technology, 2-17-1 Tsudanuma, Narashino, Chiba 275-0016, Japan.
Organizational Affiliation: