Conformational and thermodynamic changes of the repressor/DNA operator complex upon monomerization shed new light on regulation mechanisms of bacterial resistance against beta-lactam antibiotics.
Boudet, J., Duval, V., Van Melckebeke, H., Blackledge, M., Amoroso, A., Joris, B., Simorre, J.P.(2007) Nucleic Acids Res 35: 4384-4395
- PubMed: 17576674
- DOI: https://doi.org/10.1093/nar/gkm448
- Primary Citation of Related Structures:
2P7C - PubMed Abstract:
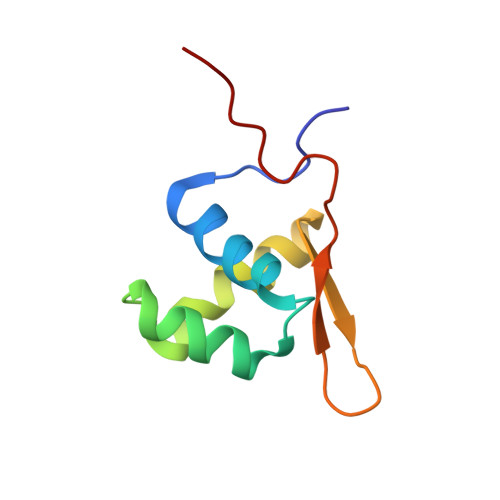
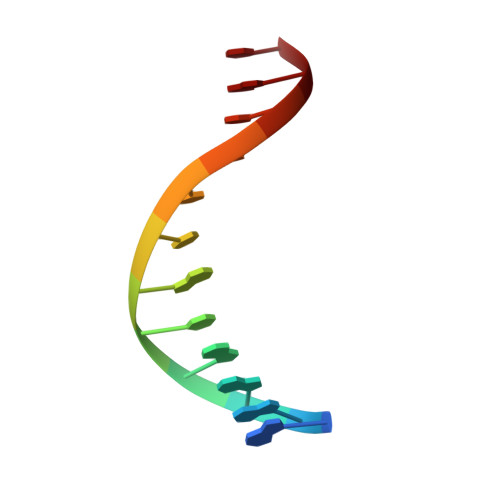
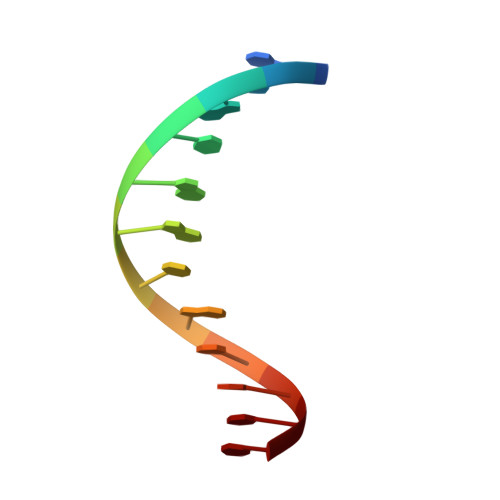
In absence of beta-lactam antibiotics, BlaI and MecI homodimeric repressors negatively control the expression of genes involved in beta-lactam resistance in Bacillus licheniformis and in Staphylococcus aureus. Subsequently to beta-lactam presence, BlaI/MecI is inactivated by a single-point proteolysis that separates its N-terminal DNA-binding domain to its C-terminal domain responsible for its dimerization. Concomitantly to this proteolysis, the truncated repressor acquires a low affinity for its DNA target that explains the expression of the structural gene for resistance. To understand the loss of the high DNA affinity of the truncated repressor, we have determined the different dissociation constants of the system and solved the solution structure of the B. licheniformis monomeric repressor complexed to the semi-operating sequence OP1 of blaP (1/2OP1blaP) by using a de novo docking approach based on inter-molecular nuclear Overhauser effects and chemical-shift differences measured on each macromolecular partner. Although the N-terminal domain of the repressor is not subject to internal structural rearrangements upon DNA binding, the molecules adopt a tertiary conformation different from the crystallographic operator-repressor dimer complex, leading to a 30 degrees rotation of the monomer with respect to a central axis extended across the DNA. These results open new insights for the repression and induction mechanisms of bacterial resistance to beta-lactams.
Organizational Affiliation:
Institut de Biologie Structurale Jean-Pierre Ebel CEA-CNRS-UJF, 41 Avenue Jules Horowitz, 38027 Grenoble Cedex 1, France.