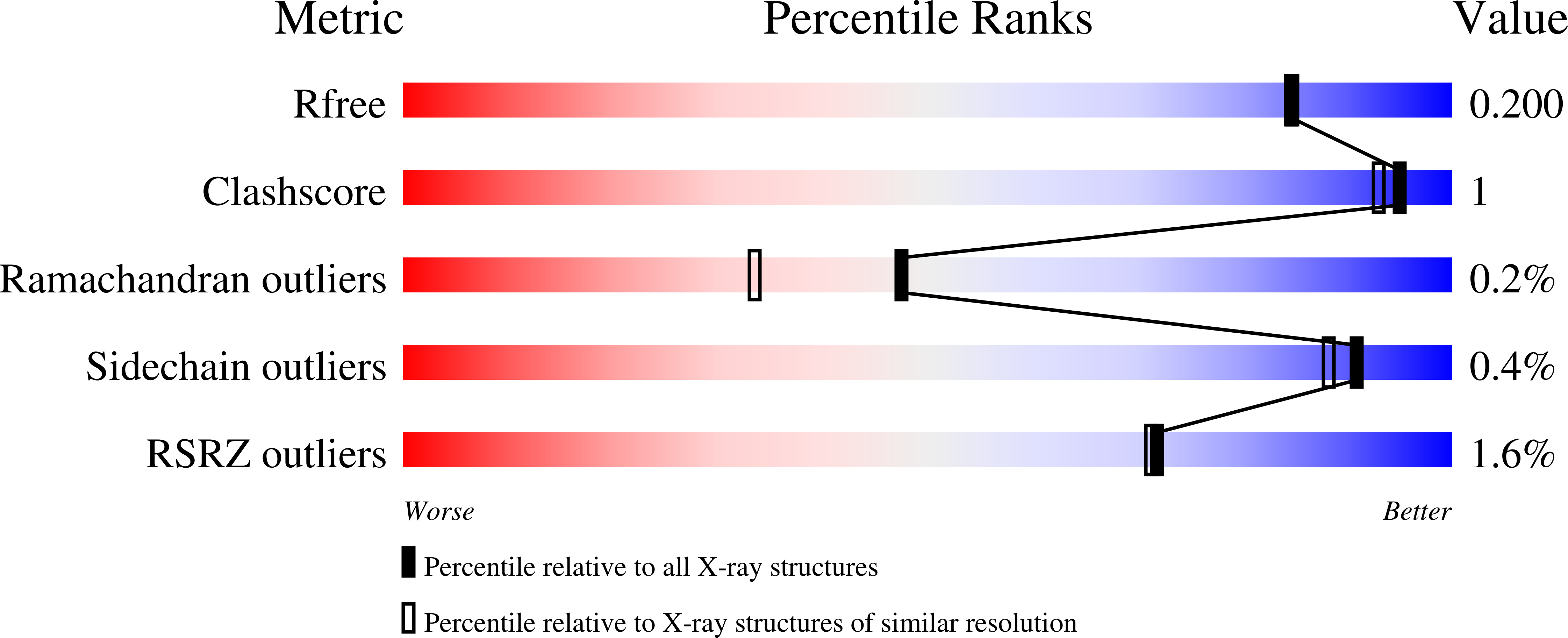
Enhanced Fructose Oxidase Activity in a Galactose Oxidase Variant
Deacon, S.E., Mahmoud, K., Spooner, R.K., Firbank, S.J., Knowles, P.F., Phillips, S.E.V., McPherson, M.J.(2004) Chembiochem 5: 972
- PubMed: 15239055
- DOI: https://doi.org/10.1002/cbic.200300810
- Primary Citation of Related Structures:
2JKX - PubMed Abstract:
Galactose oxidase (GO; EC 1.1.3.9) catalyses the oxidation of a wide range of primary alcohols including mono-, oligo- and polysaccharides. High-resolution structures have been determined for GO, but no structural information is available for the enzyme with bound substrate or inhibitor. Previously, computer-aided docking experiments have been used to develop a plausible model for interactions between GO and the D-galactose substrate. Residues implicated in such interactions include Arg330, Gln406, Phe464, Phe194 and Trp290. In the present study we describe an improved expression system for recombinant GO in the methylotrophic yeast Pichia pastoris. We use this system to express variant proteins mutated at Arg330 and Phe464 to explore the substrate binding model. We also demonstrate that the Arg330 variants display greater fructose oxidase activity than does wild-type GO.
Organizational Affiliation:
Astbury Centre for Structural Molecular Biology, School of Biochemistry and Molecular Biology, University of Leeds, Leeds, LS2 9JT, UK.