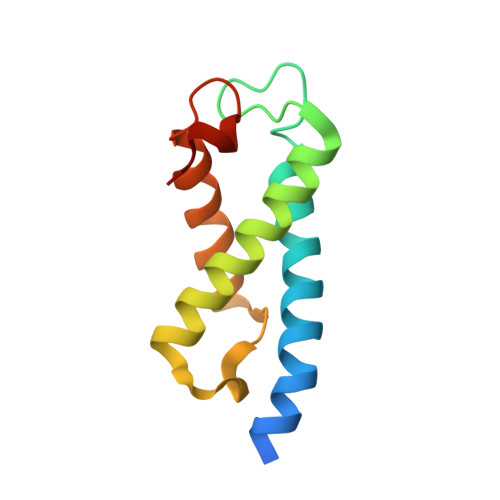
Solution structure of the Set2 Rpb1 interacting domain of human Set2 and its interaction with the hyperphosphorylated C-terminal domain of Rpb1
Li, M., Phatnani, H.P., Guan, Z., Sage, H., Greenleaf, A., Zhou, P.(2005) Proc Natl Acad Sci U S A 102: 17636-17641
- PubMed: 16314571
- DOI: https://doi.org/10.1073/pnas.0506350102
- Primary Citation of Related Structures:
2A7O - PubMed Abstract:
The phosphorylation state of the C-terminal repeat domain (CTD) of the largest subunit of RNA polymerase II changes as polymerase transcribes a gene, and the distinct forms of the phospho-CTD (PCTD) recruit different nuclear factors to elongating polymerase. The Set2 histone methyltransferase from yeast was recently shown to bind the PCTD of elongating RNA polymerase II by means of a novel domain termed the Set2-Rpb1 interacting (SRI) domain. Here, we report the solution structure of the SRI domain in human Set2 (hSRI domain), which adopts a left-turned three-helix bundle distinctly different from other structurally characterized PCTD-interacting domains. NMR titration experiments mapped the binding surface of the hSRI domain to helices 1 and 2, and Biacore binding studies showed that the domain binds preferably to [Ser-2 + Ser-5]-phosphorylated CTD peptides containing two or more heptad repeats. Point-mutagenesis studies identified five residues critical for PCTD binding. In view of the differential effects of these point mutations on binding to different CTD phosphopeptides, we propose a model for the hSRI domain interaction with the PCTD.
Organizational Affiliation:
Department of Biochemistry, Duke University Medical Center, Research Drive, Durham, NC 27710, USA.