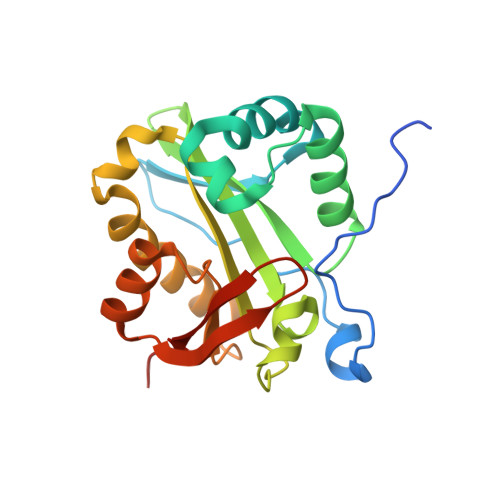
The crystal structure of Rv1347c, a putative antibiotic resistance protein from Mycobacterium tuberculosis, reveals a GCN5-related fold and suggests an alternative function in siderophore biosynthesis
Card, G.L., Peterson, N.A., Smith, C.A., Rupp, B., Schick, B.M., Baker, E.N.(2005) J Biol Chem 280: 13978-13986
- PubMed: 15695811
- DOI: https://doi.org/10.1074/jbc.M413904200
- Primary Citation of Related Structures:
1YK3 - PubMed Abstract:
Mycobacterium tuberculosis, the cause of tuberculosis, is a devastating human pathogen. The emergence of multidrug resistance in recent years has prompted a search for new drug targets and for a better understanding of mechanisms of resistance. Here we focus on the gene product of an open reading frame from M. tuberculosis, Rv1347c, which is annotated as a putative aminoglycoside N-acetyltransferase. The Rv1347c protein does not show this activity, however, and we show from its crystal structure, coupled with functional and bioinformatic data, that its most likely role is in the biosynthesis of mycobactin, the M. tuberculosis siderophore. The crystal structure of Rv1347c was determined by multiwavelength anomalous diffraction phasing from selenomethionine-substituted protein and refined at 2.2 angstrom resolution (r = 0.227, R(free) = 0.257). The protein is monomeric, with a fold that places it in the GCN5-related N-acetyltransferase (GNAT) family of acyltransferases. Features of the structure are an acyl-CoA binding site that is shared with other GNAT family members and an adjacent hydrophobic channel leading to the surface that could accommodate long-chain acyl groups. Modeling the postulated substrate, the N(epsilon)-hydroxylysine side chain of mycobactin, into the acceptor substrate binding groove identifies two residues at the active site, His130 and Asp168, that have putative roles in substrate binding and catalysis.
Organizational Affiliation:
School of Biological Sciences, University of Auckland, Auckland, New Zealand.