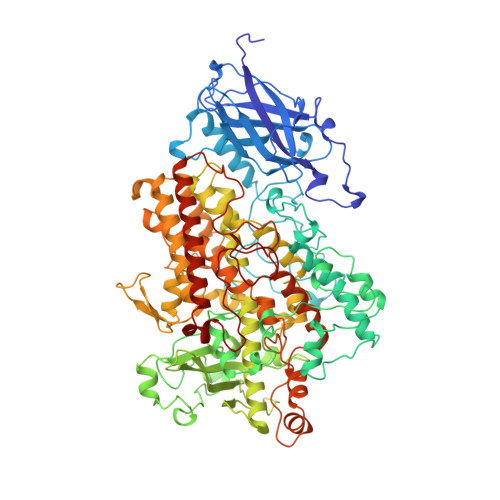
Crystal structure of soybean lipoxygenase L-1 at 1.4 A resolution.
Minor, W., Steczko, J., Stec, B., Otwinowski, Z., Bolin, J.T., Walter, R., Axelrod, B.(1996) Biochemistry 35: 10687-10701
- PubMed: 8718858
- DOI: https://doi.org/10.1021/bi960576u
- Primary Citation of Related Structures:
1YGE - PubMed Abstract:
Lipoxygenases, which are widely distributed among plant and animal species, are Fe-containing dioxygenases that act on lipids containing (Z,Z)-pentadiene moieties in the synthesis of compounds with a variety of functions. Utilizing an improved strategy of data collection, low temperature, and synchrotron radiation of short wavelength, the structure of ferrous soybean lipoxygenase L-1, a single chain protein of 839 amino acid residues, has been determined by X-ray crystallography to a resolution of 1.4 A. The R-factor for the refined model is 19.7%. General features of the protein structure were found to be consistent with the results of prior crystallographic studies at lower (2.6 A) resolution. In contrast to the prior studies, the binding of a water molecule to the active site Fe was established. The octahedral coordination sphere of the Fe also includes the side chains of His499, His504, His690, and Asn694 as well as the terminal carboxylate of Ile839, which binds as a monodentate ligand. Asn694 is involved in a number of labile polar interactions with other protein groups, including an amide-aromatic hydrogen bond, and appears to be a weak ligand. Several possible access routes for dioxygen and fatty acids to the internal active site and substrate binding cavity are described. The protein structure restricts access to the Fe site such that the formation of an organo-Fe intermediate seems improbable. Structural restrictions pertinent to other proposed reaction intermediates, such as planar pentadienyl and nonplanar allyl radicals, are also discussed.
Organizational Affiliation:
Department of Biological Sciences, Purdue University, West Lafayette, Indiana 47907, USA.