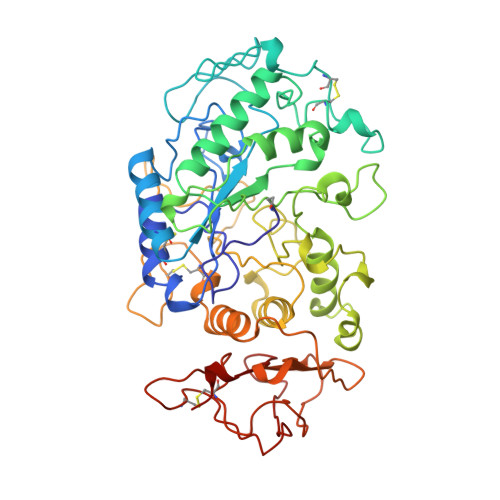
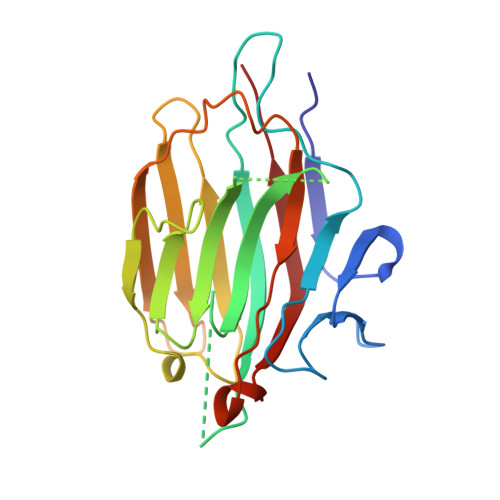
A plant-seed inhibitor of two classes of alpha-amylases: X-ray analysis of Tenebrio molitor larvae alpha-amylase in complex with the bean Phaseolus vulgaris inhibitor.
Nahoum, V., Farisei, F., Le-Berre-Anton, V., Egloff, M.P., Rouge, P., Poerio, E., Payan, F.(1999) Acta Crystallogr D Biol Crystallogr 55: 360-362
- PubMed: 10089450
- DOI: https://doi.org/10.1107/S0907444998010701
- Primary Citation of Related Structures:
1VIW - PubMed Abstract:
The alpha-amylase from Tenebrio molitor larvae (TMA) has been crystallized in complex with the alpha-amylase inhibitor (alpha-AI) from the bean Phaseolus vulgaris. A molecular-replacement solution of the structure was obtained using the refined pig pancreatic alpha-amylase (PPA) and alpha-AI atomic coordinates as starting models. The structural analysis showed that although TMA has the typical structure common to alpha-amylases, large deviations from the mammalian alpha-amylase models occur in the loops. Despite these differences in the interacting loops, the bean inhibitor is still able to inhibit both the insect and mammalian alpha-amylase.
Organizational Affiliation:
AFMB-IBSM-CNRS, 31 Chemin Joseph Aiguier, 13402 Marseille CEDEX 20, France.