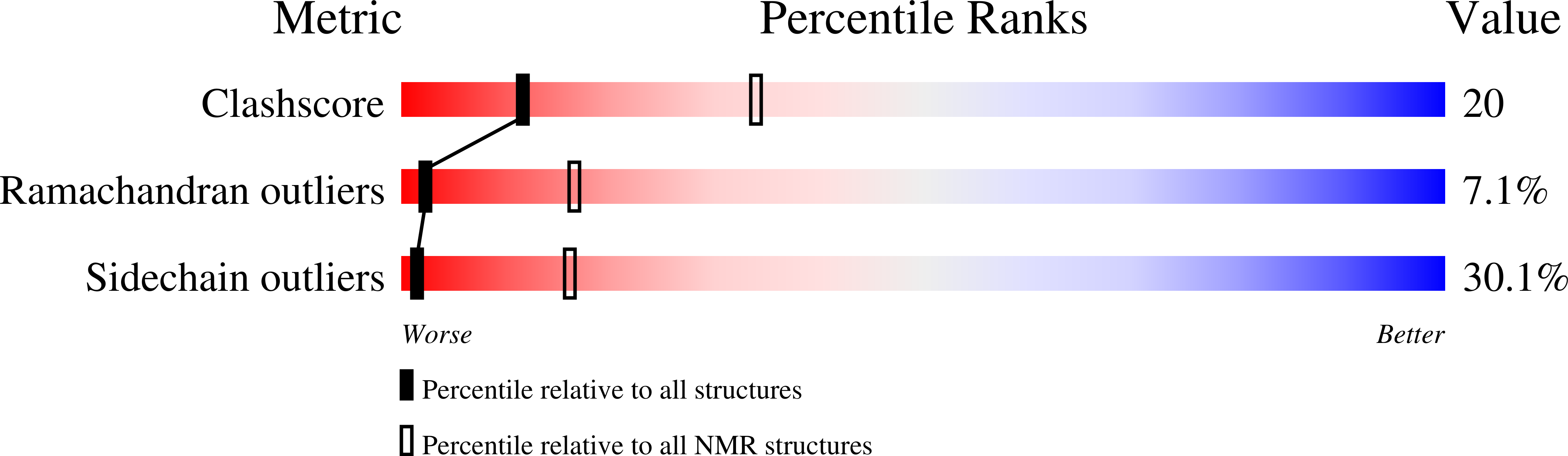Solution structure of the flexible class II ubiquitin-conjugating enzyme Ubc1 provides insights for polyubiquitin chain assembly.
Merkley, N., Shaw, G.S.(2004) J Biol Chem 279: 47139-47147
- PubMed: 15328341
- DOI: https://doi.org/10.1074/jbc.M409576200
- Primary Citation of Related Structures:
1TTE - PubMed Abstract:
E2 conjugating enzymes form a thiol ester intermediate with ubiquitin, which is subsequently transferred to a substrate protein targeted for degradation. While all E2 proteins comprise a catalytic domain where the thiol ester is formed, several E2s (class II) have C-terminal extensions proposed to control substrate recognition, dimerization, or polyubiquitin chain formation. Here we present the novel solution structure of the class II E2 conjugating enzyme Ubc1 from Saccharomyces cerevisiae. The structure shows the N-terminal catalytic domain adopts an alpha/beta fold typical of other E2 proteins. This domain is physically separated from its C-terminal domain by a 22-residue flexible tether. The C-terminal domain adopts a three-helix bundle that we have identified as an ubiquitin-associated domain (UBA). NMR chemical shift perturbation experiments show this UBA domain interacts in a regioselective manner with ubiquitin. This two-domain structure of Ubc1 was used to identify other UBA-containing class II E2 proteins, including human E2-25K, that likely have a similar architecture and to determine the role of the UBA domain in facilitating polyubiquitin chain formation.
Organizational Affiliation:
Department of Biochemistry, The University of Western Ontario, London, Ontario N6A 5C1, Canada.