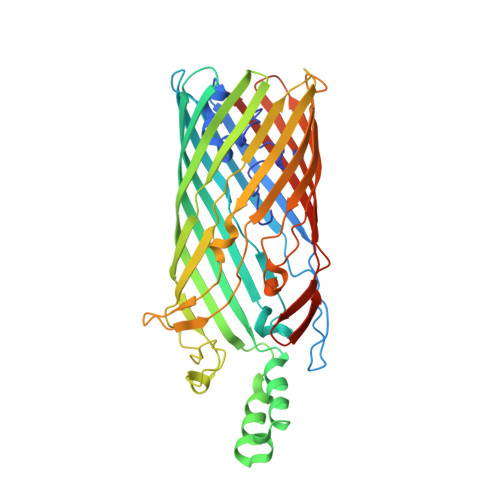
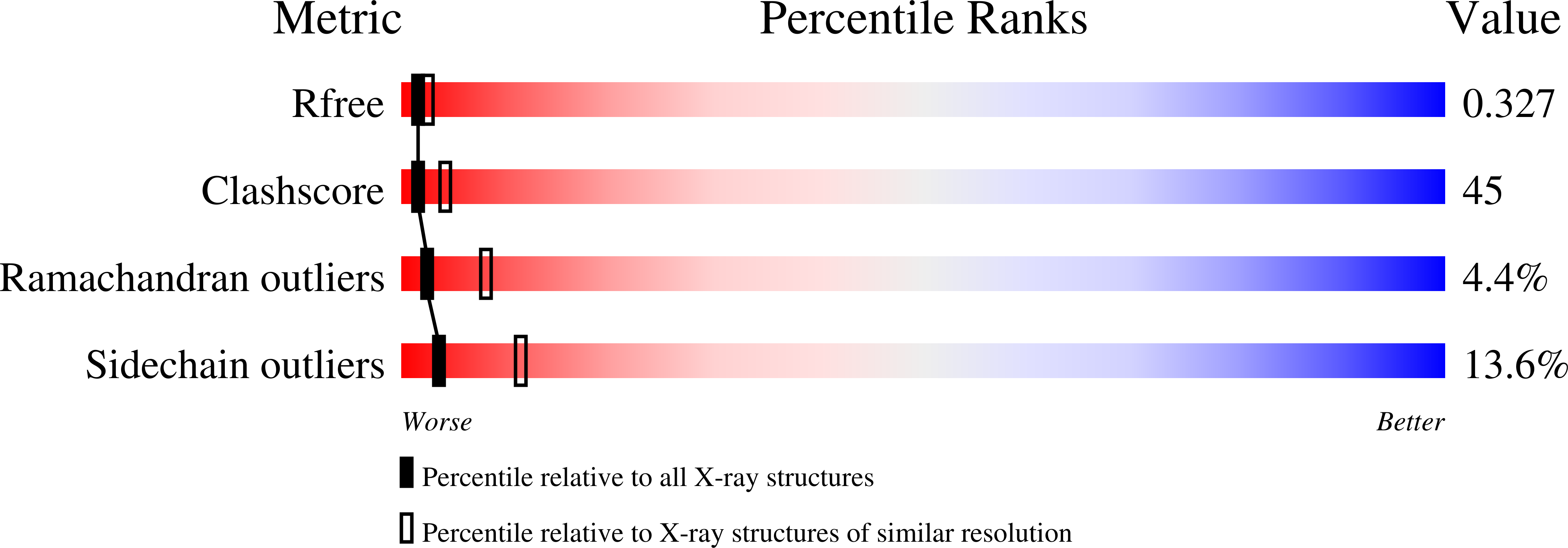Crystal structure of the long-chain fatty acid transporter FadL.
van den Berg, B., Black, P.N., Clemons Jr., W.M., Rapoport, T.A.(2004) Science 304: 1506-1509
- PubMed: 15178802
- DOI: https://doi.org/10.1126/science.1097524
- Primary Citation of Related Structures:
1T16, 1T1L - PubMed Abstract:
The mechanisms by which hydrophobic molecules, such as long-chain fatty acids, enter cells are poorly understood. In Gram-negative bacteria, the lipopolysaccharide layer in the outer membrane is an efficient barrier for fatty acids and aromatic hydrocarbons destined for biodegradation. We report crystal structures of the long-chain fatty acid transporter FadL from Escherichia coli at 2.6 and 2.8 angstrom resolution. FadL forms a 14-stranded beta barrel that is occluded by a central hatch domain. The structures suggest that hydrophobic compounds bind to multiple sites in FadL and use a transport mechanism that involves spontaneous conformational changes in the hatch.
Organizational Affiliation:
Howard Hughes Medical Institute and Department of Cell Biology, Harvard Medical School, 240 Longwood Avenue, Boston, MA 02115, USA. lvandenberg@hms.harvard.edu