Very fast folding and association of a trimerization domain from bacteriophage t4 fibritin.
Guthe, S., Kapinos, L., Moglich, A., Meier, S., Grzesiek, S., Kiefhaber, T.(2004) J Mol Biol 337: 905-915
- PubMed: 15033360
- DOI: https://doi.org/10.1016/j.jmb.2004.02.020
- Primary Citation of Related Structures:
1RFO - PubMed Abstract:
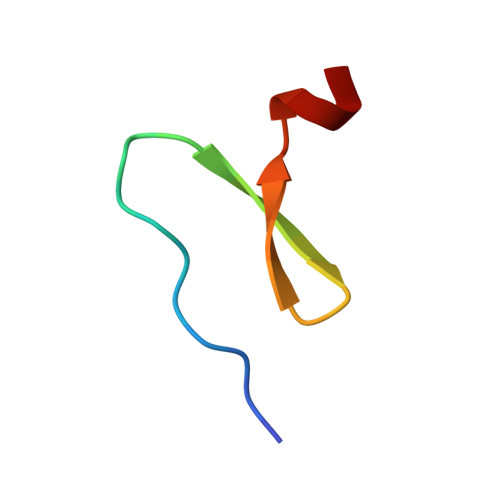
The foldon domain constitutes the C-terminal 30 amino acid residues of the trimeric protein fibritin from bacteriophage T4. Its function is to promote folding and trimerization of fibritin. We investigated structure, stability and folding mechanism of the isolated foldon domain. The domain folds into the same trimeric beta-propeller structure as in fibritin and undergoes a two-state unfolding transition from folded trimer to unfolded monomers. The folding kinetics involve several consecutive reactions. Structure formation in the region of the single beta-hairpin of each monomer occurs on the submillisecond timescale. This reaction is followed by two consecutive association steps with rate constants of 1.9(+/-0.5)x10(6)M(-1)s(-1) and 5.4(+/-0.3)x10(6)M(-1)s(-1) at 0.58 M GdmCl, respectively. This is similar to the fastest reported bimolecular association reactions for folding of dimeric proteins. At low concentrations of protein, folding shows apparent third-order kinetics. At high concentrations of protein, the reaction becomes almost independent of protein concentrations with a half-time of about 3 ms, indicating that a first-order folding step from a partially folded trimer to the native protein (k=210 +/- 20 s(-1)) becomes rate-limiting. Our results suggest that all steps on the folding/trimerization pathway of the foldon domain are evolutionarily optimized for rapid and specific initiation of trimer formation during fibritin assembly. The results further show that beta-hairpins allow efficient and rapid protein-protein interactions during folding.
Organizational Affiliation:
Division of Biophysical Chemistry, Biozentrum der Universität Basel, Klingelbergstrasse 70, CH-4056 Basel, Switzerland.