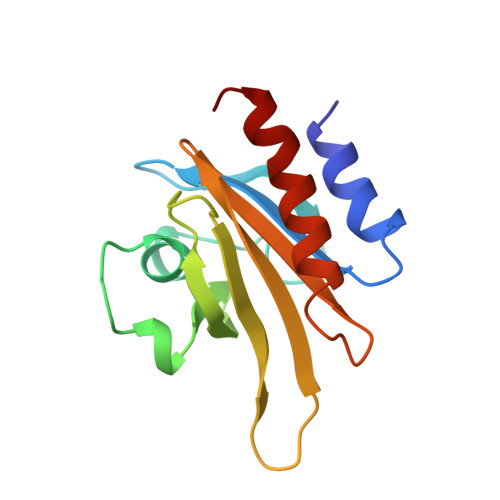
Refined solution structure of human profilin I.
Metzler, W.J., Farmer 2nd., B.T., Constantine, K.L., Friedrichs, M.S., Lavoie, T., Mueller, L.(1995) Protein Sci 4: 450-459
- PubMed: 7795529
- DOI: https://doi.org/10.1002/pro.5560040312
- Primary Citation of Related Structures:
1PFL - PubMed Abstract:
Profilin is a ubiquitous eukaryotic protein that binds to both cytosolic actin and the phospholipid phosphatidylinositol-4,5-bisphosphate. These dual competitive binding capabilities of profilin suggest that profilin serves as a link between the phosphatidyl inositol cycle and actin polymerization, and thus profilin may be an essential component in the signaling pathway leading to cytoskeletal rearrangement. The refined three-dimensional solution structure of human profilin I has been determined using multidimensional heteronuclear NMR spectroscopy. Twenty structures were selected to represent the solution conformational ensemble. This ensemble of structures has root-mean-square distance deviations from the mean structure of 0.58 A for the backbone atoms and 0.98 A for all non-hydrogen atoms. Comparison of the solution structure of human profilin to the crystal structure of bovine profilin reveals that, although profilin adopts essentially identical conformations in both states, the solution structure is more compact than the crystal structure. Interestingly, the regions that show the most structural diversity are located at or near the actin-binding site of profilin. We suggest that structural differences are reflective of dynamical properties of profilin that facilitate favorable interactions with actin. The global folding pattern of human profilin also closely resembles that of Acanthamoeba profilin I, reflective of the 22% sequence identity and approximately 45% sequence similarity between these two proteins.
Organizational Affiliation:
Department of Macromolecular NMR, Bristol-Myers Squibb Pharmaceutical Research Institute, Princeton, New Jersey 08543-400, USA.