Distinct Requirements for Heparin and Alpha-Dystroglycan Binding Revealed by Structure-Based Mutagenesis of the Laminin Alpha2 Lg4-Lg5 Domain Pair
Wizemann, H., Garbe, J.H.O., Friedrich, M.V.K., Timpl, R., Sasaki, T., Hohenester, E.(2003) J Mol Biol 332: 635
- PubMed: 12963372
- DOI: https://doi.org/10.1016/s0022-2836(03)00848-9
- Primary Citation of Related Structures:
1OKQ - PubMed Abstract:
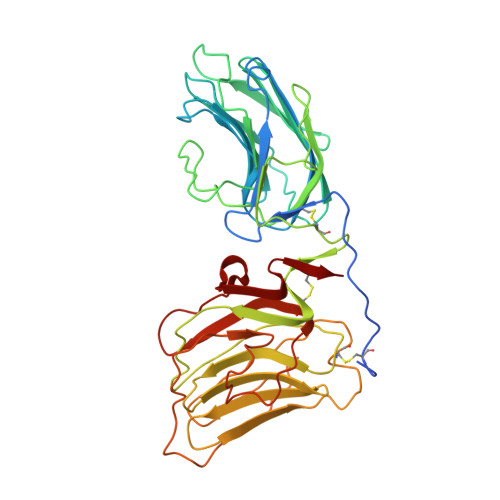
Laminin-2 (alpha2beta1gamma1) is found in basement membranes surrounding muscle and peripheral nerve cells. Several types of cellular receptors bind to the laminin G-like (LG) domains at the C terminus of the alpha2 chain, the interaction with alpha-dystroglycan (alpha-DG) being particularly important in muscle. We have used site-directed mutagenesis and in vitro binding assays to map the binding sites on the laminin alpha2 chain LG4-LG5 domain pair for alpha-DG, heparin and sulfatides. Calcium-dependent alpha-DG recognition requires the calcium ion in LG4, but not the one in LG5, as well as basic residues in both LG domains. Heparin and sulfatides also bind to basic residues in both LG domains, but there is little overlap in the binding sites for alpha-DG and heparin/sulfatides. The results should prove useful for the molecular dissection of laminin-receptor interactions in vivo.
Organizational Affiliation:
Max-Planck-Institut für Biochemie, D-82152, Martinsried, Germany.