NMR-based Binding Screen and Structural Analysis of the Complex Formed between alpha-Cobratoxin and an 18-mer Cognate Peptide Derived from the alpha1 Subunit of the Nicotinic Acetylcholine Receptor from Torpedo californica
Zeng, H., Hawrot, E.(2002) J Biol Chem 277: 37439-37445
- PubMed: 12133834
- DOI: https://doi.org/10.1074/jbc.M205483200
- Primary Citation of Related Structures:
1LXG, 1LXH - PubMed Abstract:
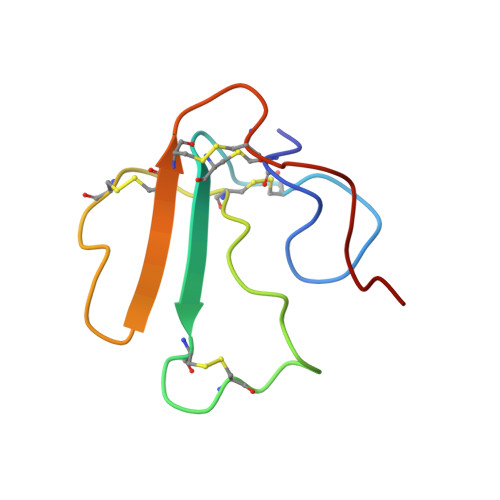
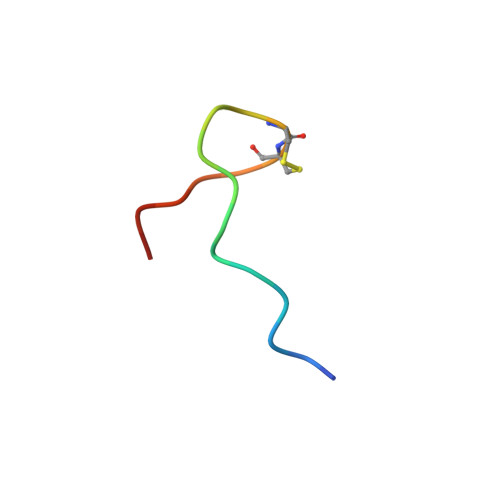
The alpha18-mer peptide, spanning residues 181-198 of the Torpedo nicotinic acetylcholine receptor alpha1 subunit, contains key binding determinants for agonists and competitive antagonists. To investigate whether the alpha18-mer can bind other alpha-neurotoxins besides alpha-bungarotoxin, we designed a two-dimensional (1)H-(15)N heteronuclear single quantum correlation experiment to screen four related neurotoxins for their binding ability to the peptide. Of the four toxins tested (erabutoxin a, erabutoxin b, LSIII, and alpha-cobratoxin), only alpha-cobratoxin binds the alpha18-mer to form a 1:1 complex. The NMR solution structure of the alpha-cobratoxin.alpha18-mer complex was determined with a backbone root mean square deviation of 1.46 A. In the structure, alpha-cobratoxin contacts the alpha18-mer at the tips of loop I and II and through C-terminal cationic residues. The contact zone derived from the intermolecular nuclear Overhauser effects is in agreement with recent biochemical data. Furthermore, the structural models support the involvement of cation-pi interactions in stabilizing the complex. In addition, the binding screen results suggest that C-terminal cationic residues of alpha-bungarotoxin and alpha-cobratoxin contribute significantly to binding of the alpha18-mer. Finally, we present a structural model for nicotinic acetylcholine receptor-alpha-cobratoxin interaction by superimposing the alpha-cobratoxin.alpha18-mer complex onto the crystal structure of the acetylcholine-binding protein (Protein Data Bank code ).
Organizational Affiliation:
Department of Molecular Pharmacology, Physiology, and Biotechnology, Brown Medical School, Providence, Rhode Island 02912, USA.