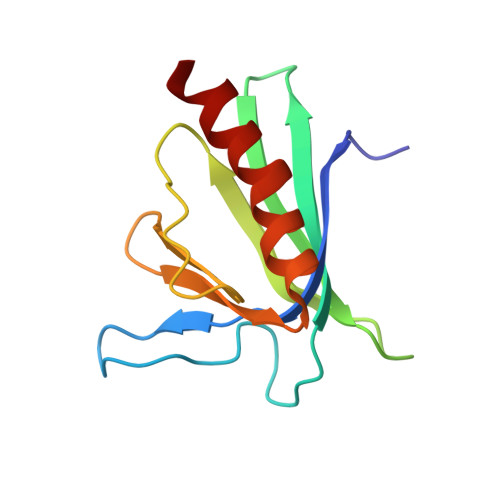
Dual epitope recognition by the VASP EVH1 domain modulates polyproline ligand specificity and binding affinity.
Ball, L.J., Kuhne, R., Hoffmann, B., Hafner, A., Schmieder, P., Volkmer-Engert, R., Hof, M., Wahl, M., Schneider-Mergener, J., Walter, U., Oschkinat, H., Jarchau, T.(2000) EMBO J 19: 4903-4914
- PubMed: 10990454
- DOI: https://doi.org/10.1093/emboj/19.18.4903
- Primary Citation of Related Structures:
1EGX - PubMed Abstract:
The Ena-VASP family of proteins act as molecular adaptors linking the cytoskeletal system to signal transduction pathways. Their N-terminal EVH1 domains use groups of exposed aromatic residues to specifically recognize 'FPPPP' motifs found in the mammalian zyxin and vinculin proteins, and ActA protein of the intracellular bacterium Listeria monocytogenes. Here, evidence is provided that the affinities of these EVH1-peptide interactions are strongly dependent on the recognition of residues flanking the core FPPPP motifs. Determination of the VASP EVH1 domain solution structure, together with peptide library screening, measurement of individual K(d)s by fluorescence titration, and NMR chemical shift mapping, revealed a second affinity-determining epitope present in all four ActA EVH1-binding motifs. The epitope was shown to interact with a complementary hydrophobic site on the EVH1 surface and to increase strongly the affinity of ActA for EVH1 domains. We propose that this epitope, which is absent in the sequences of the native EVH1-interaction partners zyxin and vinculin, may provide the pathogen with an advantage when competing for the recruitment of the host VASP and Mena proteins in the infected cell.
Organizational Affiliation:
Forschungsinstitut für Molekulare Pharmakologie, Alfred-Kowalke Strasse 4, D-10315, Berlin, Germany.