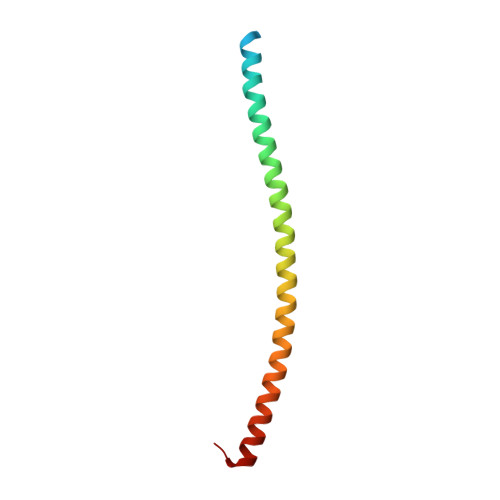Visualization of an unstable coiled coil from the scallop myosin rod
Li, Y., Brown, J.H., Reshetnikova, L., Blazsek, A., Farkas, L., Nyitray, L., Cohen, C.(2003) Nature 424: 341-345
- PubMed: 12867988
- DOI: https://doi.org/10.1038/nature01801
- Primary Citation of Related Structures:
1NKN - PubMed Abstract:
Alpha-helical coiled coils in muscle exemplify simplicity and economy of protein design: small variations in sequence lead to remarkable diversity in cellular functions. Myosin II is the key protein in muscle contraction, and the molecule's two-chain alpha-helical coiled-coil rod region--towards the carboxy terminus of the heavy chain--has unusual structural and dynamic features. The amino-terminal subfragment-2 (S2) domains of the rods can swing out from the thick filament backbone at a hinge in the coiled coil, allowing the two myosin 'heads' and their motor domains to interact with actin and generate tension. Most of the S2 rod appears to be a flexible coiled coil, but studies suggest that the structure at the N-terminal region is unstable, and unwinding or bending of the alpha-helices near the head-rod junction seems necessary for many of myosin's functional properties. Here we show the physical basis of a particularly weak coiled-coil segment by determining the 2.5-A-resolution crystal structure of a leucine-zipper-stabilized fragment of the scallop striated-muscle myosin rod adjacent to the head-rod junction. The N-terminal 14 residues are poorly ordered; the rest of the S2 segment forms a flexible coiled coil with poorly packed core residues. The unusual absence of interhelical salt bridges here exposes apolar core atoms to solvent.
- Rosenstiel Basic Medical Sciences Research Center, Brandeis University, Waltham, Massachusetts 02454-9110, USA.
Organizational Affiliation: