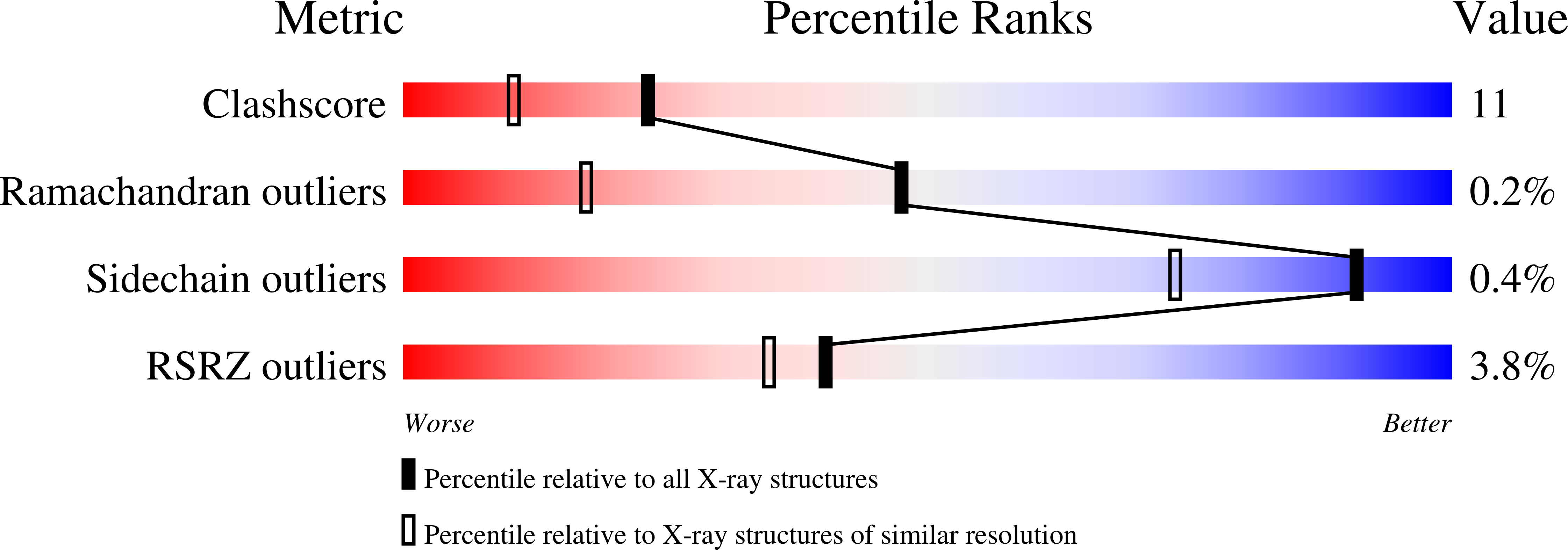A covalent modification of NADP+ revealed by the atomic resolution structure of FprA, a Mycobacterium tuberculosis oxidoreductase.
Bossi, R.T., Aliverti, A., Raimondi, D., Fischer, F., Zanetti, G., Ferrari, D., Tahallah, N., Maier, C.S., Heck, A.J.R., Rizzi, M., Mattevi, A.(2002) Biochemistry 41: 8807-8818
- PubMed: 12102623
- DOI: https://doi.org/10.1021/bi025858a
- Primary Citation of Related Structures:
1LQT, 1LQU - PubMed Abstract:
FprA is a mycobacterial oxidoreductase that catalyzes the transfer of reducing equivalents from NADPH to a protein acceptor. We determined the atomic resolution structure of FprA in the oxidized (1.05 A resolution) and NADPH-reduced (1.25 A resolution) forms. The comparison of these FprA structures with that of bovine adrenodoxin reductase showed no significant overall differences. Hence, these enzymes, which belong to the structural family of the disulfide oxidoreductases, are structurally conserved in very distant organisms such as mycobacteria and mammals. Despite the conservation of the overall fold, the details of the active site of FprA show some peculiar features. In the oxidized enzyme complex, the bound NADP+ exhibits a covalent modification, which has been identified as an oxygen atom linked through a carbonylic bond to the reactive C4 atom of the nicotinamide ring. Mass spectrometry has confirmed this assignment. This NADP+ derivative is likely to form by oxidation of the NADP+ adduct resulting from nucleophilic attack by an active-site water molecule. A Glu-His pair is well positioned to activate the attacking water through a mechanism analogous to that of the catalytic triad in serine proteases. The NADP+ nicotinamide ring exhibits the unusual cis conformation, which may favor derivative formation. The physiological significance of this reaction is presently unknown. However, it could assist with drug-design studies in that the modified NADP+ could serve as a lead compound for the development of specific inhibitors.
Organizational Affiliation:
Dipartimento di Genetica e Microbiologia, Università di Pavia, via Abbiategrasso 207, 27100 Pavia, Italy.