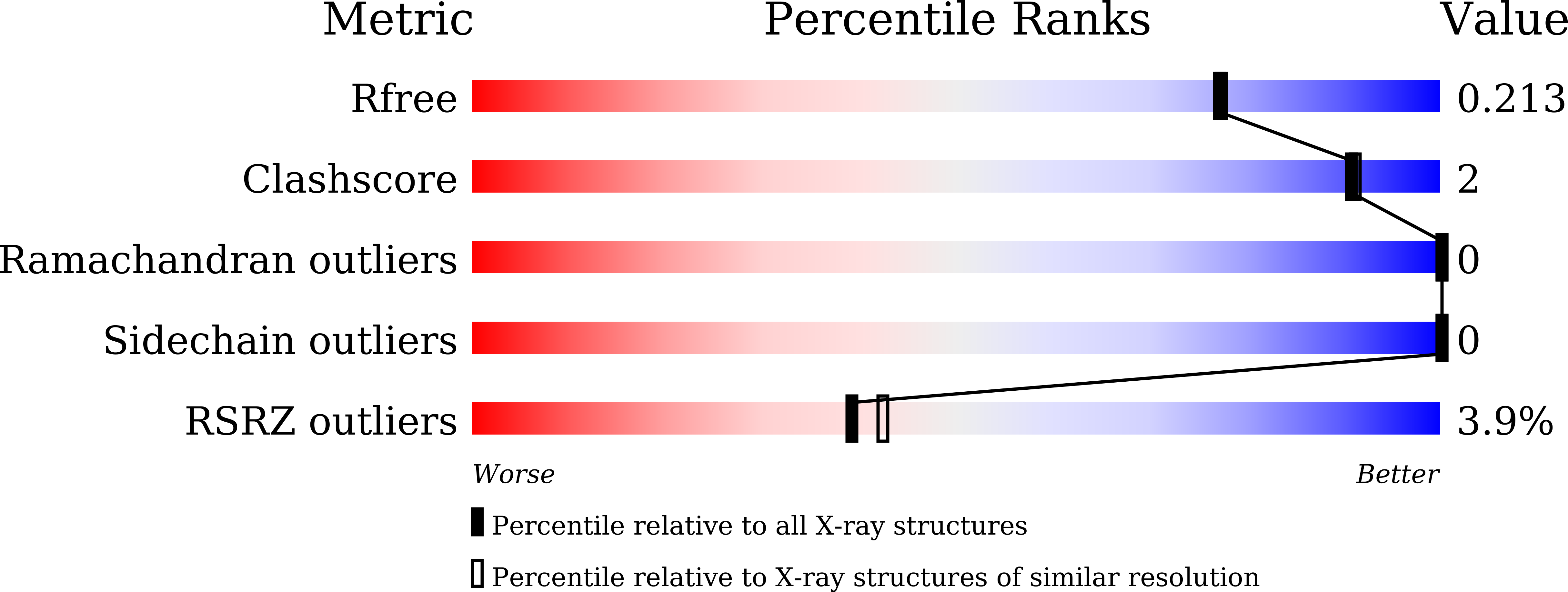
Structure-based development and preclinical evaluation of the SARS-CoV-2 3C-like protease inhibitor simnotrelvir.
Jiang, X., Su, H., Shang, W., Zhou, F., Zhang, Y., Zhao, W., Zhang, Q., Xie, H., Jiang, L., Nie, T., Yang, F., Xiong, M., Huang, X., Li, M., Chen, P., Peng, S., Xiao, G., Jiang, H., Tang, R., Zhang, L., Shen, J., Xu, Y.(2023) Nat Commun 14: 6463-6463
- PubMed: 37833261
- DOI: https://doi.org/10.1038/s41467-023-42102-y
- Primary Citation of Related Structures:
8IFP, 8IFQ, 8IFR, 8IFS, 8IFT, 8IGX, 8IGY - PubMed Abstract:
The persistent pandemic of coronavirus disease 2019 (COVID-19) caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and its variants accentuates the great demand for developing effective therapeutic agents. Here, we report the development of an orally bioavailable SARS-CoV-2 3C-like protease (3CL pro ) inhibitor, namely simnotrelvir, and its preclinical evaluation, which lay the foundation for clinical trials studies as well as the conditional approval of simnotrelvir in combination with ritonavir for the treatment of COVID-19. The structure-based optimization of boceprevir, an approved HCV protease inhibitor, leads to identification of simnotrelvir that covalently inhibits SARS-CoV-2 3CL pro with an enthalpy-driven thermodynamic binding signature. Multiple enzymatic assays reveal that simnotrelvir is a potent pan-CoV 3CL pro inhibitor but has high selectivity. It effectively blocks replications of SARS-CoV-2 variants in cell-based assays and exhibits good pharmacokinetic and safety profiles in male and female rats and monkeys, leading to robust oral efficacy in a male mouse model of SARS-CoV-2 Delta infection in which it not only significantly reduces lung viral loads but also eliminates the virus from brains. The discovery of simnotrelvir thereby highlights the utility of structure-based development of marked protease inhibitors for providing a small molecule therapeutic effectively combatting human coronaviruses.
Organizational Affiliation:
State Key Laboratory of Drug Research, Shanghai Institute of Materia Medica, Chinese Academy of Sciences, 201203, Shanghai, China.