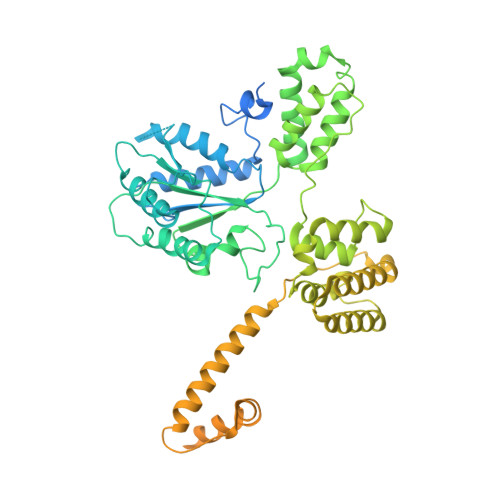
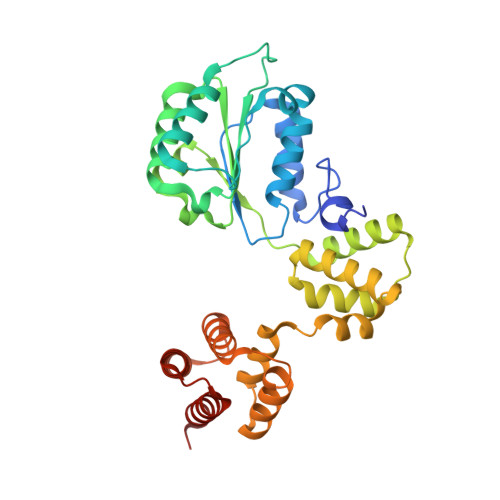
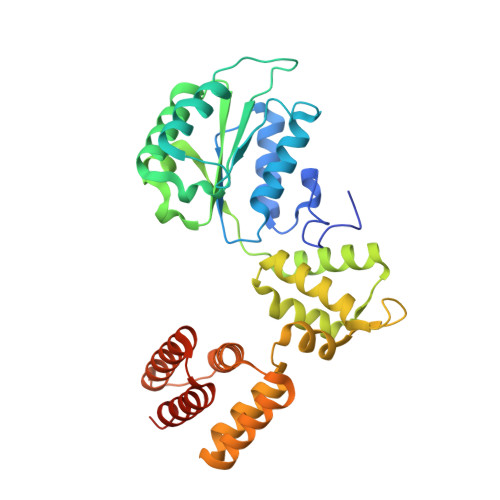
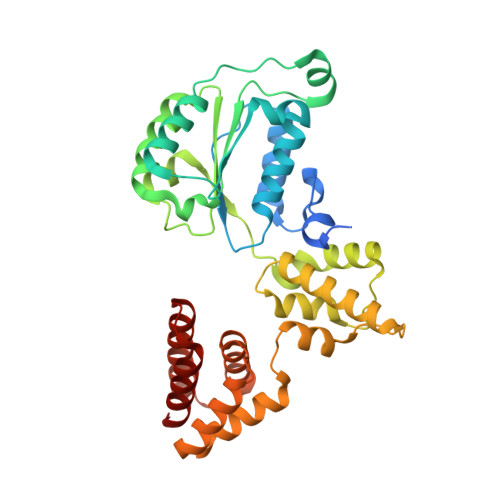
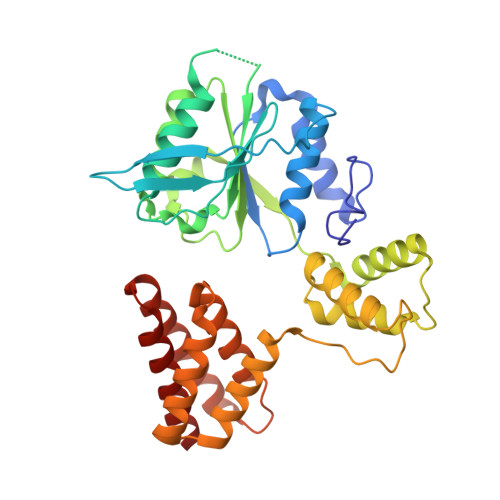
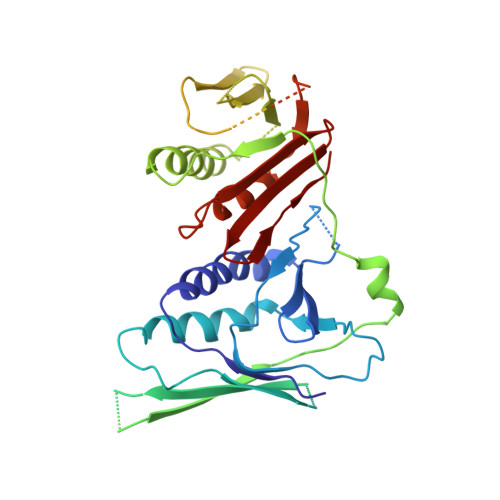
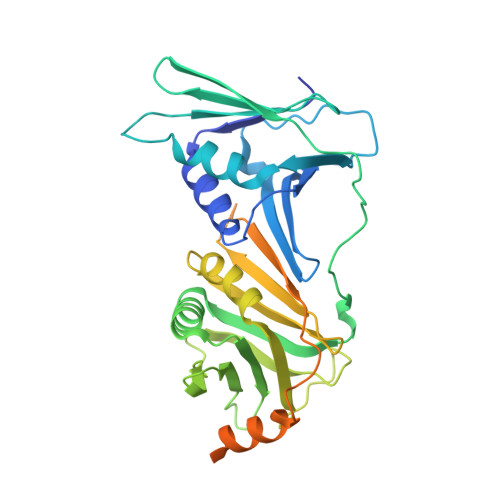
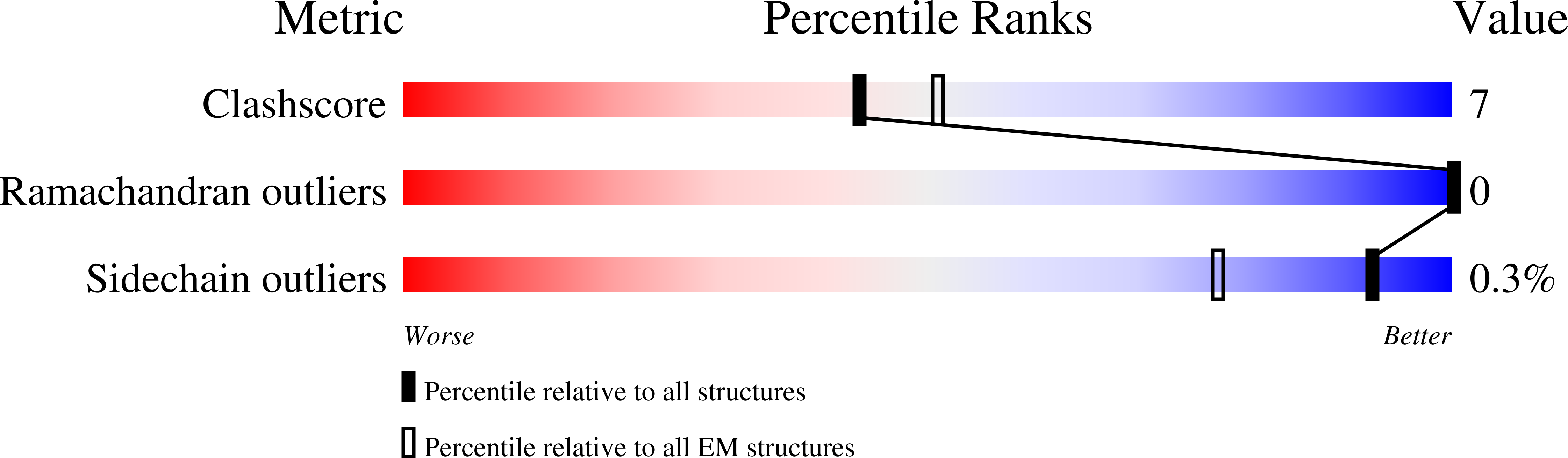DNA is loaded through the 9-1-1 DNA checkpoint clamp in the opposite direction of the PCNA clamp.
Zheng, F., Georgescu, R.E., Yao, N.Y., O'Donnell, M.E., Li, H.(2022) Nat Struct Mol Biol 29: 376-385
- PubMed: 35314830
- DOI: https://doi.org/10.1038/s41594-022-00742-6
- Primary Citation of Related Structures:
7SGZ, 7SH2 - PubMed Abstract:
The 9-1-1 DNA checkpoint clamp is loaded onto 5'-recessed DNA to activate the DNA damage checkpoint that arrests the cell cycle. The 9-1-1 clamp is a heterotrimeric ring that is loaded in Saccharomyces cerevisiae by Rad24-RFC (hRAD17-RFC), an alternate clamp loader in which Rad24 replaces Rfc1 in the RFC1-5 clamp loader of proliferating cell nuclear antigen (PCNA). The 9-1-1 clamp loading mechanism has been a mystery, because, unlike RFC, which loads PCNA onto a 3'-recessed junction, Rad24-RFC loads the 9-1-1 ring onto a 5'-recessed DNA junction. Here we report two cryo-EM structures of Rad24-RFC-DNA with a closed or 27-Å open 9-1-1 clamp. The structures reveal a completely unexpected mechanism by which a clamp can be loaded onto DNA. Unlike RFC, which encircles DNA, Rad24 binds 5'-DNA on its surface, not inside the loader, and threads the 3' ssDNA overhang into the 9-1-1 clamp from above the ring.
Organizational Affiliation:
Department of Structural Biology, Van Andel Institute, Grand Rapids, MI, USA.